Andrew J Hill
@ahill_tweets
Scientist at @infinimmune, cat enthusiast. Formerly @tunetx_news, @10xgenomics, @JShendure and @coletrapnell labs, @dgmacarthur lab.
ID: 1972974192
http://www.andrewjohnhill.com 19-10-2013 22:21:04
667 Tweet
524 Followers
249 Following


Looking forward to co-moderating a great set of talks with the wonderful @mollygasp. Join us and our exciting lineup of speakers: Jesse Engreitz, Wendy Bickmore, F. Hormozdiari, and E. Weeks today (Wed) at 3:30ET for our session on variant-to-function coupling! #ASHG20


First foray into single cell genomics, first postdoc paper, first tweet! Excited to share our human cell atlas of fetal chromatin accessibility, a very fun collaboration with Jay Shendure Andrew J Hill Riza Daza Darren Cusanovich and others. science.sciencemag.org/content/370/65… (1/5)

We are looking for a research assistant to join our lab Columbia BME. Our lab develops and applies single-cell genomic tools to understand how cancer responds to therapy. Join us! To apply: opportunities.columbia.edu/en-us/job/5127…

We’re hiring a molbio RA in Seattle! You'd be the newest member of our bad ass in vitro team nested in CajalNeuro's Functional Genomics group. You'd use your cloning skills for neurodegen high throughput projects (spoiler- largely learning new CRISPR screen skills from me! :).



We are recruting postdoctoral fellows to join us in defining responses to exposure using single cell genomics. We will review postdoc applications in January. DM, email ([email protected]) or use the link below to apply!
Well fellas, I tried the 10x Genomics’ RNA Fix solution…and it was a breeze. Not surprising easy to follow protocols, descriptions of what to expect were spot on, and there were zero challenging steps…even when I did use crazy challenging samples! Sequencing will follow 🔜

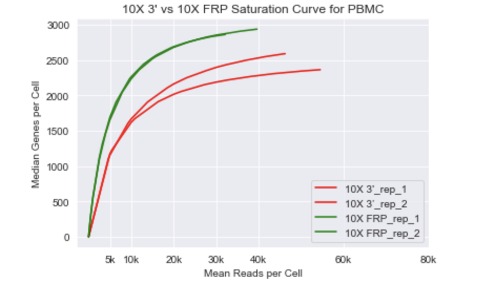
We wondered what a side-to-side comparison between 3’ v3.1 & RNA Fix solution from 10x Genomics would look like? Well, we have WINNER! I said it two #10Xperiences ago…this kit is by far the best these guys have ever produced yet! Andres F Vallejo 🧬 💻 Jens Durruthy Dr. Jasmine Plummer 👩🏻🔬🧬🇨🇦🇺🇸 Geoffrey McDermott



Our lab's latest protocol, more brilliant work from @bethkarenmartin & Chengxiang Qiu We can't send to bioRxiv b/c they desk-reject all protocols ;) Onwards to eLife - the journal where I predict Michael Eisen's assessment will be "delicious" Non-paywalled PDF: tinyurl.com/3zsyabpp




Excited to share our preprint biorxiv.org/content/10.110… (my first)! This was an amazing collaboration with the incredibly talented and rare gem Jean-Benoît (JB) Lalanne. Also indebted to a rockstar team Silvia Domcke Diego Calderon (diegoisworking@bluesky) @bethkarenmartin Tony Li Chase Suiter Cole Trapnell! 🧵(1/10)

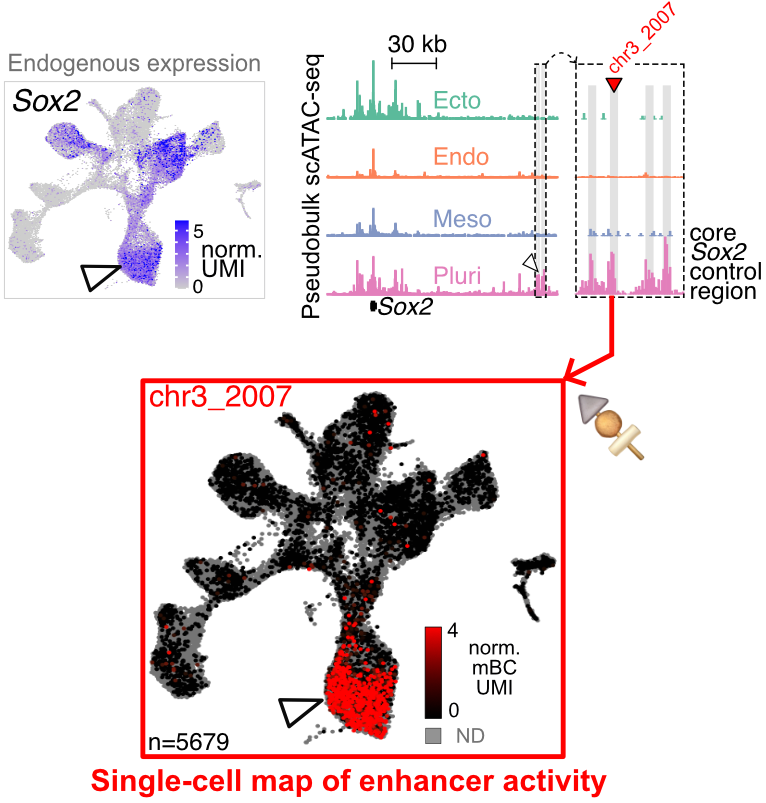
Our latest preprint from brilliant Jean-Benoît (JB) Lalanne & Samuel Gabriel Regalado is new single cell MPRA assay (sQers) with remarkable quantitative range, precision approaching Poisson shot noise. We apply sQers to discover autonomous germ layer-specific enhancers. PDF here: tinyurl.com/ym8r2jat


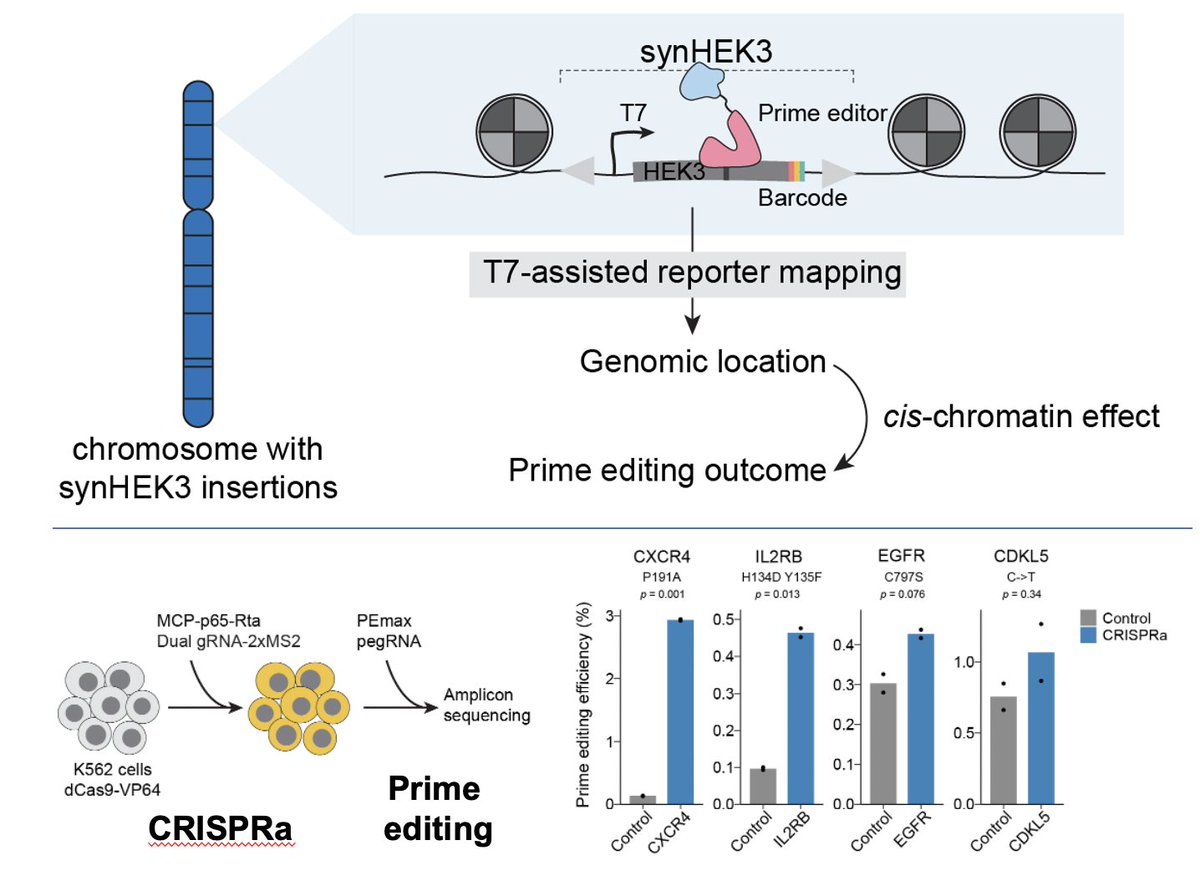
Absolutely thrilled to share mine and Troy McDiarmid’s work describing the development of a multiplexed, scalable CRISPRa cell type-specific screening framework that is able to identify…(1/n) biorxiv.org/content/10.110…