Tony Li
@tonylihq
PhD student @uwgenome
ID: 1405955662975160324
https://scholar.google.com/citations?hl=en&user=AHbq6uwAAAAJ&view_op=list_works&sortby=pubdate 18-06-2021 18:30:04
75 Tweet
61 Followers
194 Following


scQers are quantitative single cell expression reporters reported by Jay Shendure and colleagues. They enable high-sensitivity quantitative characterization of developmental cis-regulatory elements at the single-cell level. nature.com/articles/s4159…
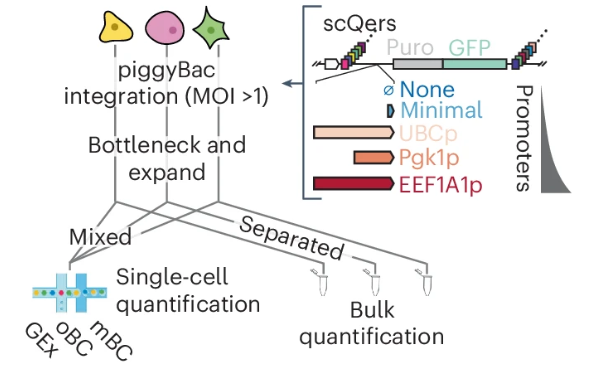


Finally out in Nature Methods from the brilliant Jean-Benoît (JB) Lalanne & Samuel Gabriel Regalado, our highly quantitive single cell MPRA (scQer), applied to mammalian embryoids to find autonomous enhancers. Bonus = Tornado circular barcodes that are all kinds of useful. OA link: rdcu.be/dHqo1
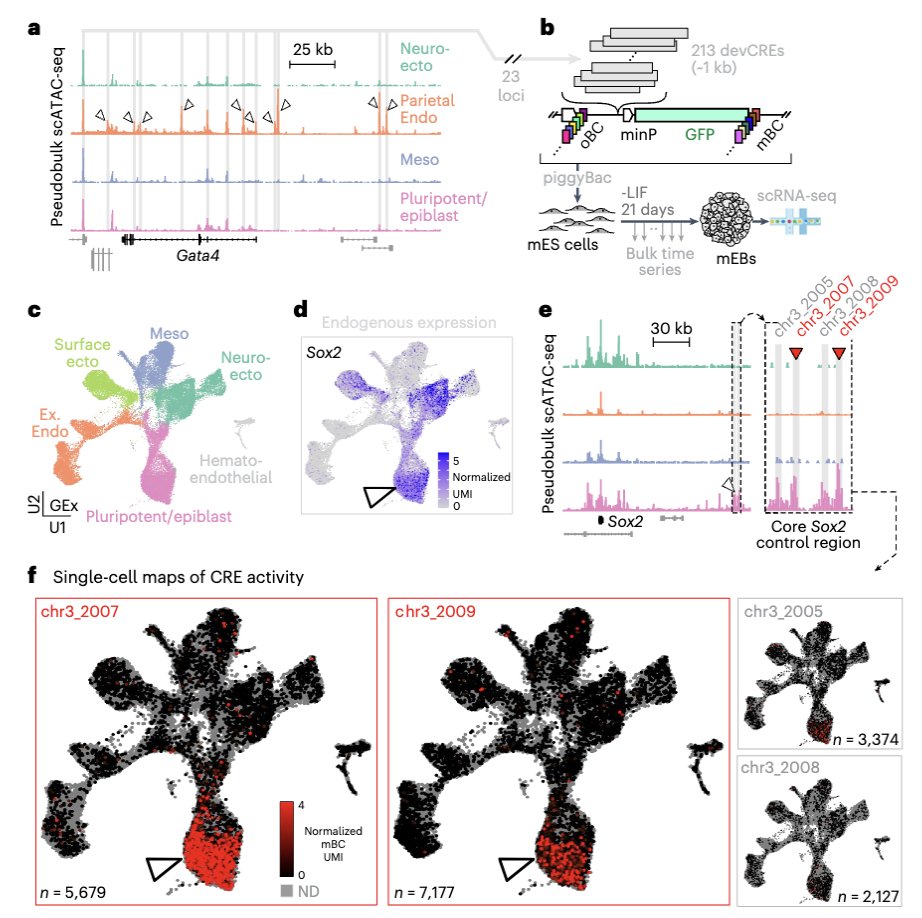


Our DNA Typewriter protocol (written by Hanna Liao, Jay Shendure, and myself) is out! Thank you Nature Protocols for writing/editing this with us and having it as this week’s #FeaturedProtocol :-)



Excited to share our human RA-gastruloid model in Nature Cell Biology! Early retinoic acid supplementation, followed by later Matrigel addition, robustly induces human embryonic morphologies and diverse cell lineages, including somitic, renal, cardiac, and neural cell types in
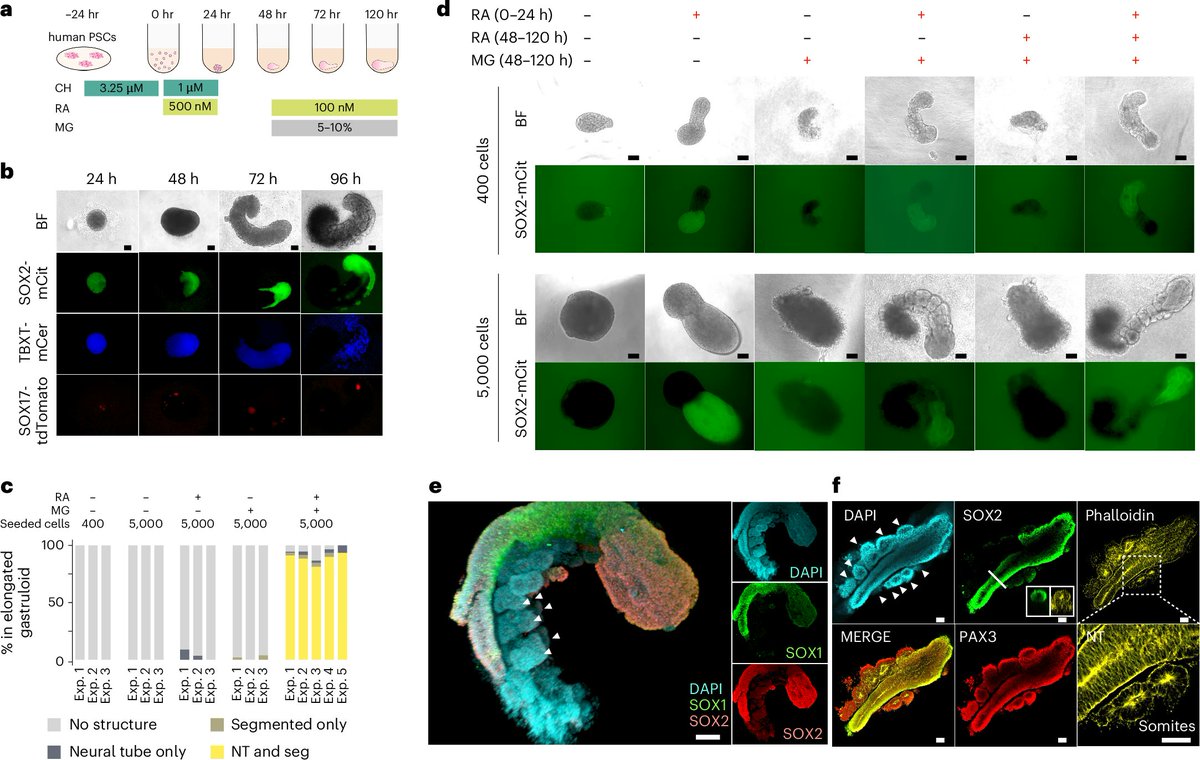

I am so thrilled to share our work of human RA gastruloid finally out on Nature Cell Biology. It has been a wonderful journey to work with a fantastic stem cell scientist Nobu Hamazaki and many talented collaborators in Jay Shendure lab. Excited to contribute to the field of stembryo!


Very happy to share our latest w/ Devin Schweppe and Nobu Hamazaki labs, where we map the temporal dynamics and proteomic landscape of mammalian gastruloids. Sneek Peak 🧵below #stembryo #gastruloid #proteomics biorxiv.org/content/10.110…


Our paper describing a scalable framework for Multiplex, single cell CRISPRa screening for cell-type specific regulatory elements is published! rdcu.be/dUnoq See the linked tweet from my awesome co-lead Flo Chardon for updates since the preprint!




Updated VISTA Enhancer Browser, 7 years in the making, out in Nucleic Acids Res! >4,500 transgenic in vivo enhancer experiments for all your developmental biology, human variation and #evodevo needs. enhancer.lbl.gov #enhancers #embryos doi.org/10.1093/nar/gk…



Now out in Science Magazine we present 'Genome-shuffle-seq': a method to shuffle mammalian genomes and characterize the impact of structural variants (SVs) with single-cell resolution in one experiment. science.org/doi/10.1126/sc…