Noorsher Ahmed
@noorsherahmed
@UCSanDiego Biomedical Sciences PhD Student | Spatial transcriptomics tech dev in @yeo_lab | @UCSDBMS | He/his
ID: 1213954814754738176
https://www.linkedin.com/in/noorsher-ahmed/ 05-01-2020 22:46:43
47 Tweet
170 Followers
297 Following





Sharing my golden birthday with Bart de Witte at #LINO23 after an absolutely amazing panel discussing the deep complexities of bringing AI to medicine!
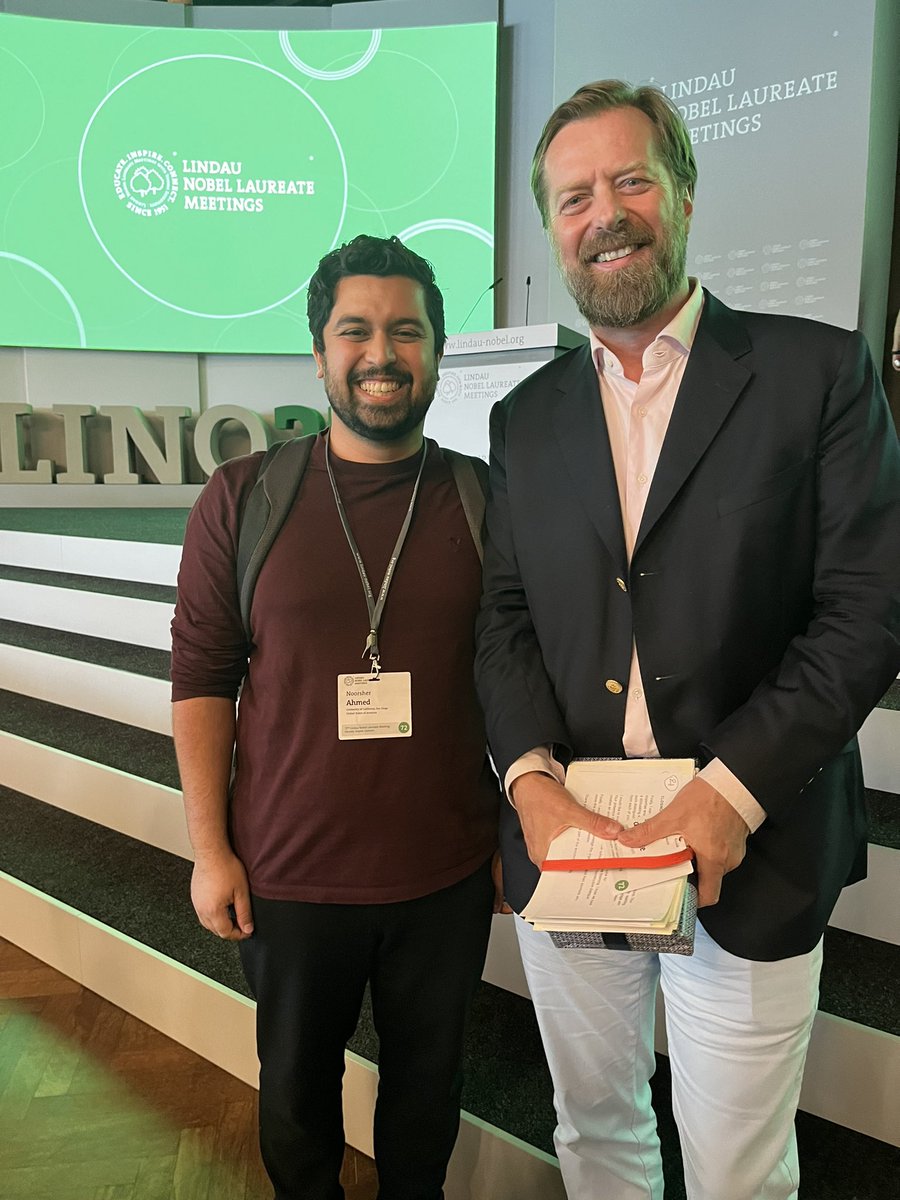



The day has come! The Chaim Lab has opened its doors UCSD BioSciences ... lab might not be ready but the incredible team is. The feeling is hard to describe. I am thankful and humbled to be where I am. Intros to lab members and lab space coming soon but here is a preview #NewPI

Brant Bassam Indeed, techs move fast. Though the true strength of #Bento lies in its universal applicability to imaging tech, rather than focusing on specific upcoming trends. This is an enduring quality that transcends time, and may well become its benchmark for assessment. Noorsher Ahmed

1/10 image-based screening advanced drug discovery but scaling to massive perturbation space is hard! Given a cell image, we asked if we could predict the morphological effect induced by a perturbation! Led by Alessandro Palma & w Fabian Theis we propose IMPA tinyurl.com/yy4jfn4h

Honored to receive the inaugural Sydney Brenner Medal to recognize the best in scholarship in the fields of life sciences, especially molecular biology not more than 20 years after the PhD. Delighted that the award was presented by Carla Brenner Academia Europaea !





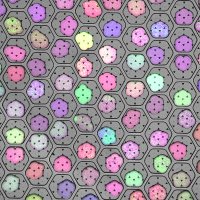
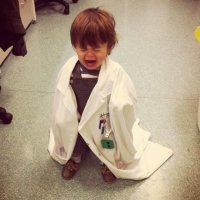
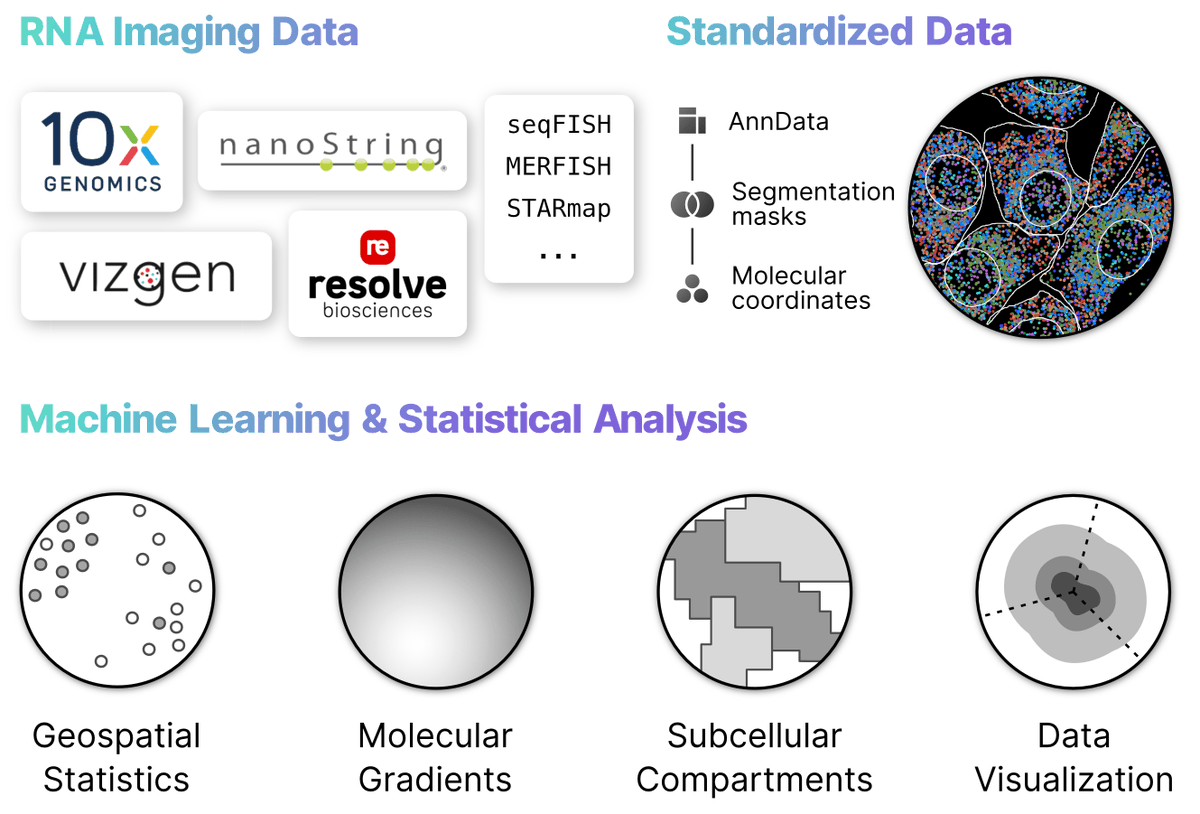
🥳 Our paper for Bento, a python toolkit for the subcellular analysis of spatial transcriptomics data is finally out in Genome Biology ! It was a privilege working with Clarence Mah @clarencemah.bsky.social and the team. Amazing that even before publication, Bento has 14k downloads, 44 Github stars!!

Announcing a huge v2.1 release for Bento, enabling even more scalable analysis of spatial transcriptomic data! 🏃💨 Plus, our paper is now published at Genome Biology after a roller coaster of preprints, submissions, revisions, new authors, and datasets 🎈 Noorsher Ahmed 1/6


Excited to share “Charlene” Hsuan-lin Her's paper describing Mudskipper software package which demonstrates superior performance over existing tools on multiplex CLIP datasets and supports analysis of repetitive elements and small non-coding RNAs Cell Genomics cell.com/cell-genomics/…

Thank you Stanford HAI and Hoffman-Yee for this grant to build a unified model of human cells spanning molecules, their structures and cellular organization - applied towards women’s health. Super excited for this collab w Stephen Quake Jure Leskovec Serena Yeung-Levy Russ Altman 🌟

7 years ago, I met a junior fellow named Jason Buenrostro who blew me away with a vision of futuristic genomic technologies Today, we (Ajay Labade, @carolinecomenho) are excited to share our first steps into that future: Expansion in situ genome sequencing 1/

Cilia and Flagella enthusiasts, stay tuned for tomorrow when the cells in the Human Protein Atlas will grow cilia and flagella. If you are at #HUPO2024, come to the plenary session tomorrow at 9.15 am by Emma Lundberg and the release of version v24 (with cilia) at 6.40 pm CET!