XiaoWang
@xiaowangzyy
ID: 792928662001549312
31-10-2016 03:18:19
40 Tweet
61 Takipçi
199 Takip Edilen

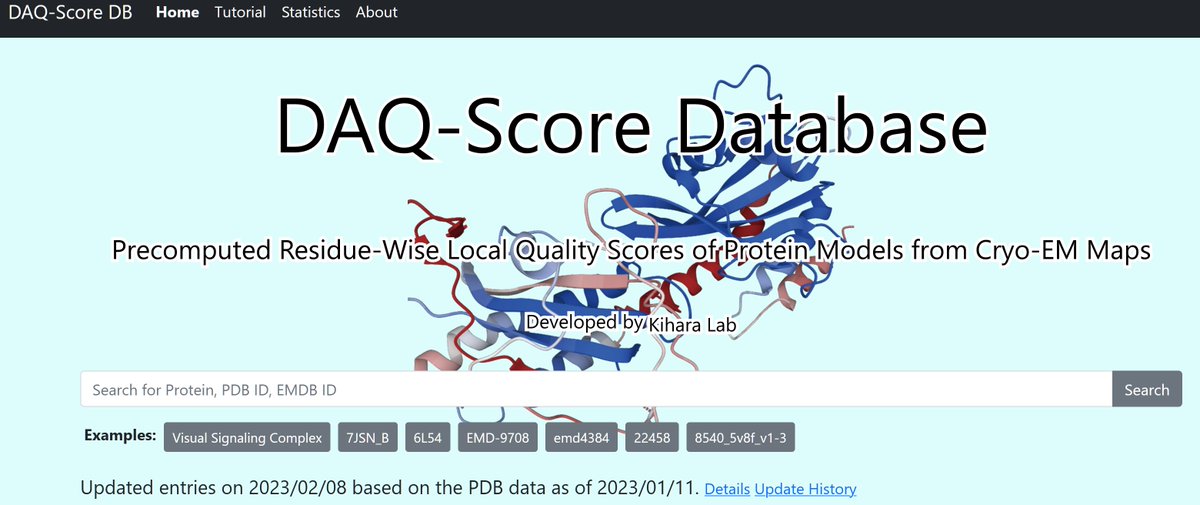
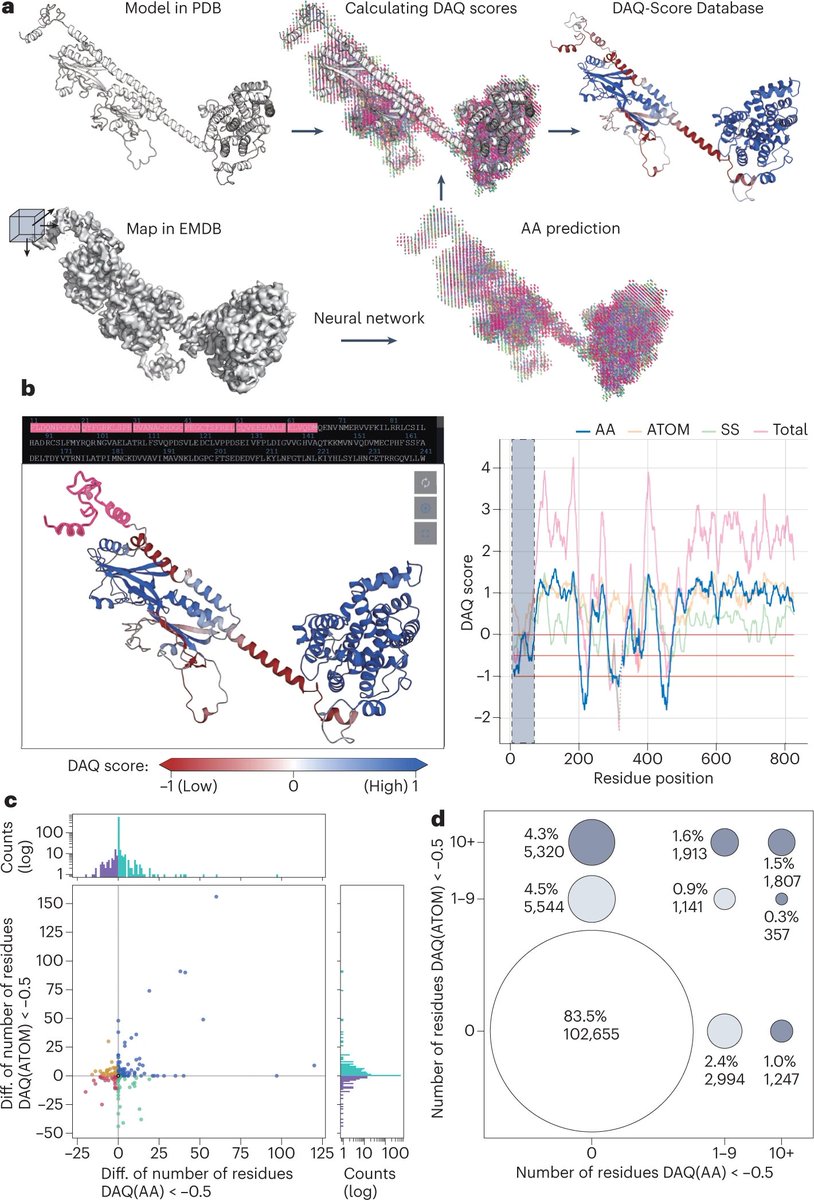
Just out online! "DAQ-Score Database: assessment of map–model compatibility for protein structure models from cryo-EM maps", Tsukasa Nakamura, Xiao Wang, Genki Terashi, & Kihara, Nature Methods Nature Methods nature.com/articles/s4159… Visit the db at daqdb.kiharalab.org


New paper from our lab! "CryoREAD: de novo structure modeling for nucleic acids in cryo-EM maps using deep learning" by @xiaowangzyy Genki Terashi Kihara, in Nature Methods nature.com/articles/s4159… Web server, G Colab, code: kiharalab.org/emsuites/cryor… @purduecs Purdue Science


Had a workshop on CryoREAD, DNA/RNA modeling tool for Cryo-EM. Discussion led by XiaoWang. It was a very well-attended event! Send us any follow-up questions. Web server: em.kiharalab.org/algorithm/Cryo… Paper: nature.com/articles/s4159… Photos: Pranav Punuru Purdue Science Purdue Computer Science

A brief introduction about our new paper CryoREAD (automated DNA/RNA structure modeling tool from cryo-EM): protocolsmethods.springernature.com/posts/cryoread…. It has been published in Nature Methods: nature.com/articles/s4159…. The free server is available at: em.kiharalab.org/algorithm/Cryo….

Behind the Paper: Blog by the first author Xiao Wang on our recent paper "CryoREAD: de novo structure modeling for nucleic acids in cryo-EM maps using deep learning" in Nature Methods. protocolsmethods.springernature.com/posts/cryoread… Server at: em.kiharalab.org/algorithm/Cryo… More at: kiharalab.org/emsuites/



My algorithm talk about CryoREAD for DNA/RNA modeling was made available on Youtube: youtube.com/watch?v=p7Bpou…. A more detailed version of our instructions in kiharalab.org/emsuites/cryor…. The paper is here: nature.com/articles/s4159…. Our online server is here: em.kiharalab.org/algorithm/Cryo….




DeepMainmast is a protein structure modeling protocol for cryo-EM that combines the strengths of a deep-learning-based de novo protein main-chain-tracing approach with AlphaFold2-based structure predictions for improved performance. Genki Terashi doi.org/10.1038/s41592…

New paper from our lab: "GO2Sum: generating human-readable functional summary of proteins from GO terms" by Swagarika Jaharlal Giri nabil_ibtehaz from Purdue Computer Science published in NPJ Systems Biology and Applications. (cont.) nature.com/articles/s4154…

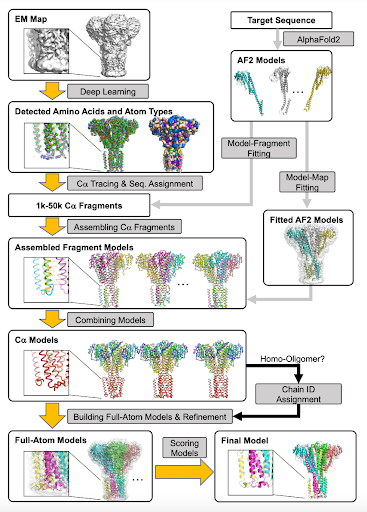
Introducing DiffModeler, a fully automated structure fitting method for modeling large protein complex structures from cryo-EM maps of a broad range of resolutions. XiaoWang Genki Terashi Daisuke Kihara Kihara Laboratory nature.com/articles/s4159…