nabil_ibtehaz
@nibtehaz
computerphile
ID: 1027556259677884416
09-08-2018 14:04:38
88 Tweet
29 Followers
76 Following

How can deep learning be used to accurately predict RNA tertiary structures when experimental determination is costly and time-consuming?Purdue University Nature Communications "NuFold: end-to-end approach for RNA tertiary structure prediction with flexible nucleobase center representation"

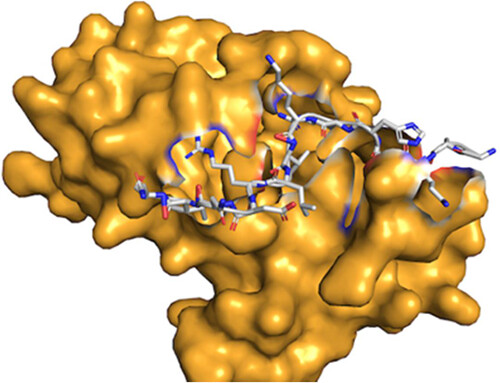
"NuFold: end-to-end approach for RNA tertiary structure prediction with flexible nucleobase center representation" by Yuki Kagaya et al. is now out in Nature Communications ! This deep learning method played a key role in our 3rd place finish for RNA in CASP16. nature.com/articles/s4146…


NuFold: end-to-end approach for RNA tertiary structure prediction with flexible nucleobase center representation Nature Communications 1. NuFold is a novel deep learning-based framework that predicts RNA tertiary structures directly from sequence data, offering a breakthrough in RNA







New paper released! "Distance-AF improves predicted protein structure models by AlphaFold2 with user-specified distance constraints" Yuanyuan Zhang, Zicong Zhang, Y Kagaya, G Terashi, B Zhao, Y Xiong & D Kihara, Communications Biology. Communications Biology nature.com/articles/s4200…