Alan Rubin
@rubin_af
Trying to figure out what genomic variants do. Computational biologist at @WEHI_research. Previously @uwgenome.
Mastodon: genomic.social/web/afrubin
ID: 3232137366
01-06-2015 03:52:47
814 Tweet
898 Followers
889 Following

Here's my review in Human Molecular Genetics on the latest and greatest multiplex assays for linking human variants to disease phenotypes. academic.oup.com/hmg/advance-ar…

@dougfowler42 and I are excited to announce the Center for Actionable Variant Analysis 🍾 has been funded by the NHGRI and the IGVF consortium. CAVA will apply multiplex assays of variant effect developed UW Genome Sciences to clinically relevant genes 1/7

Excited for our sister NHGRI Center at UW, which will focus on scaling up and applying technologies developed by our CMAP investigators @dougfowler42 Stan Fields Alan Rubin Frederick (Fritz) Roth Jay Shendure Christine Queitsch Cole Trapnell Judit Villen


Very proud (and nervous) to announce my first podcast!! We are getting scientists, doctors and comedians to talk about COVID and vaccines to help the vaccine hesitant with the info they might be looking for. Follow The Jab Gab Podcast to find out when the next first episode drops.

JD Long is now @[email protected] I'm afraid this glorious bit has not ever been recorded 😂 But I can give you a better link to use: speakerdeck.com/jennybc/how-to… (if I had known how many 👀 these slides would receive, I would made them a bit more polished)

📣 New publication from our group 📰"Closing the gap: Systematic integration of multiplexed functional data resolves variants of uncertain significance in BRCA1, TP53, and PTEN" Alan Rubin Lea Starita @dougfowler42 Shawn Fayer

Debbie Nickerson's UW Genome Sciences contributions to human genome science & medical genetics, passionate support of women & trainees in research, NHGRI funded work & "unfiltered, tenacious style" recalled by her colleagues in The New York Times obituary by Richard Sandomir nyti.ms/3nGz1ve


I'm delighted to be leading this project with such a great team. Looking forward to bringing MAVE technology into the clinic here in Australia! Shout out to all my fellow Atlas of Variant Effects Alliance members who are contributing to this mission locally and internationally.

Congratulations to Alan Rubin, Amanda Spurdle, Hamish Scott, Belinda Phipson, Paul James, @dougfowler42, Lea Starita, Chris Hahn, Anna Brown & Cliff Meldrum. WEHI (Walter and Eliza Hall Institute) Peter Mac Cancer Centre @uw Uni of Adelaide @NSWHPathology Australian Department of Health and Aged Care





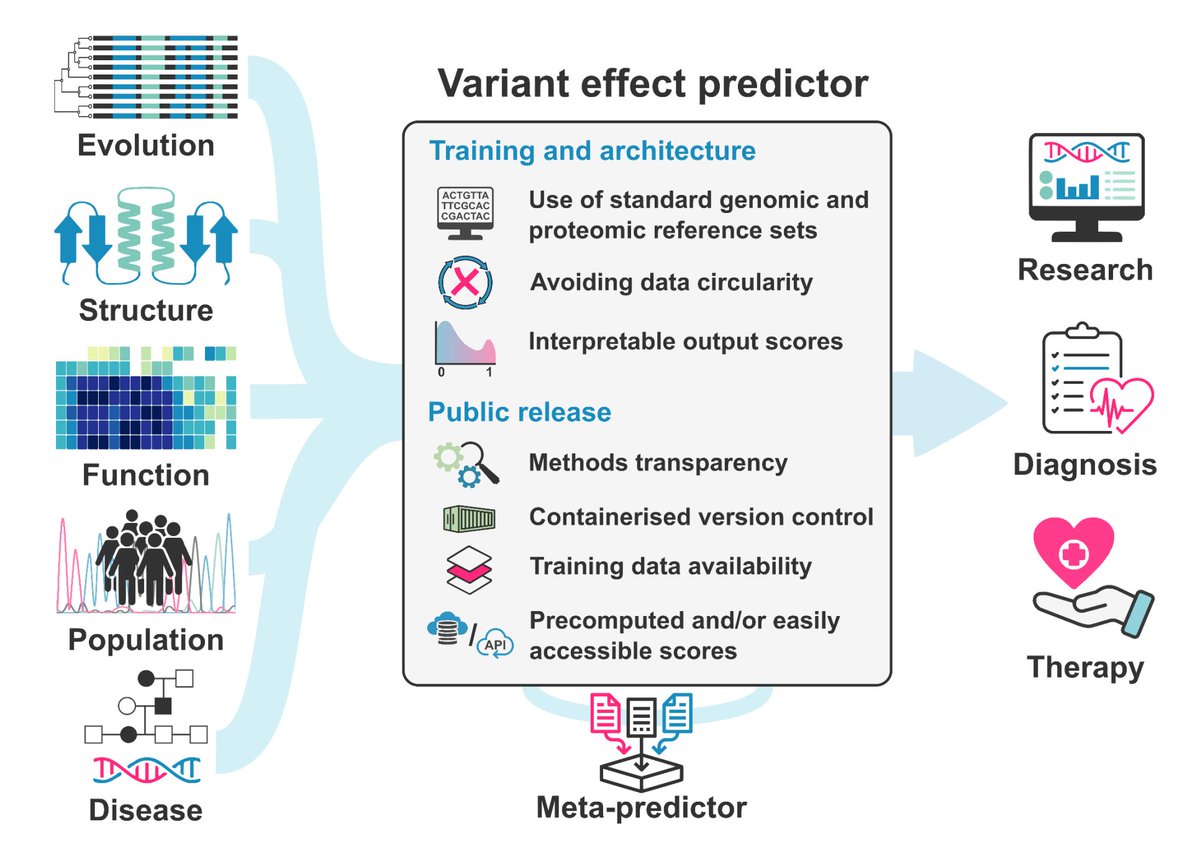
Guidelines for releasing a variant effect predictor By Ben Livesey Joe Marsh and other members of the atlas of Atlas of Variant Effects Alliance alliance 🧬 arxiv.org/abs/2404.10807

Jeremy Arbesfeld presenting pivotal work ccg2024g about a framework for mapping MAVE data, which has allowed DECIPHER and other resources to display MaveDB data ClinGen Atlas of Variant Effects Alliance Alan Rubin


Really proud of this work, the result of a fantastic international collaboration among experts brought together by Atlas of Variant Effects Alliance. Huge thanks to everyone involved!


Wonderful presentation from Eric Green (Eric Green) at Murdoch Children's Research Institute (MCRI) discussing the history of genomics and the future of genomic medicine, as part of his whirlwind tour of the Australian east coast, coordinated by Australian Genomics.



