
Eoghan Harrington
@eoghh
Current: Oxford Nanopore, Previous: Flatiron Health, 23andMe, Stanford, EMBL, TCD. Opinions my own
ID: 169634038
22-07-2010 20:21:13
224 Tweet
146 Followers
414 Following

Looking for a motivated person to join my #Algorithms and #MachineLearning team to further improve our industry leading Oxford Nanopore modified base detection capabilities!


Excited to share our paper, out today in nature, where we characterize transcriptome variation in human tissues by long-read sequencing. Huge thanks to Dafni Beryl Cummings Garrett Garborcauskas Daniel MacArthur and other collaborators. 🧵 nature.com/articles/s4158…



Minimap2 #Rust bindings update to v0.1.11. HTS lib support (SAM/BAM writing) and mm2-fast support via feature flag. Huge thanks to Eoghan Harrington for the HTS lib support! Check out the full announcement: sci.kiwi/@josephguhlin/… #bioinformatics #nanopore #genomics






#ASMicrobe folks: come see Christine He at her poster to learn how Oxford Nanopore sequencing of human microbiome samples can simultaneously: - Deplete host DNA to get more microbial reads - Generate genome-resolved assemblies - Provide host DNA methylation from the proximal tissue

Bacterial epigenomes are very cool and extremely diverse! And you get them for free when you do Oxford Nanopore sequencing 🧫🧬

Another LOADED update from the Oxford Nanopore EPI2ME team with wf-metagenomics and wf-somatic-variation getting new functionality & our desktop app getting easier for Windows users & displaying workflow progress. All this and more here: labs.epi2me.io/epi2me-23.07-0… We’re also hiring…

Here The Banfield Lab used #nanopore sequencing to study 'the nature and existence of enigmatic genetic elements' (Borgs) in soil. They assembled 7 new curated Borg genomes, revealing extensive DNA methylation that differentiates them from their host archaea, Methanoperedens. 🧵1/2

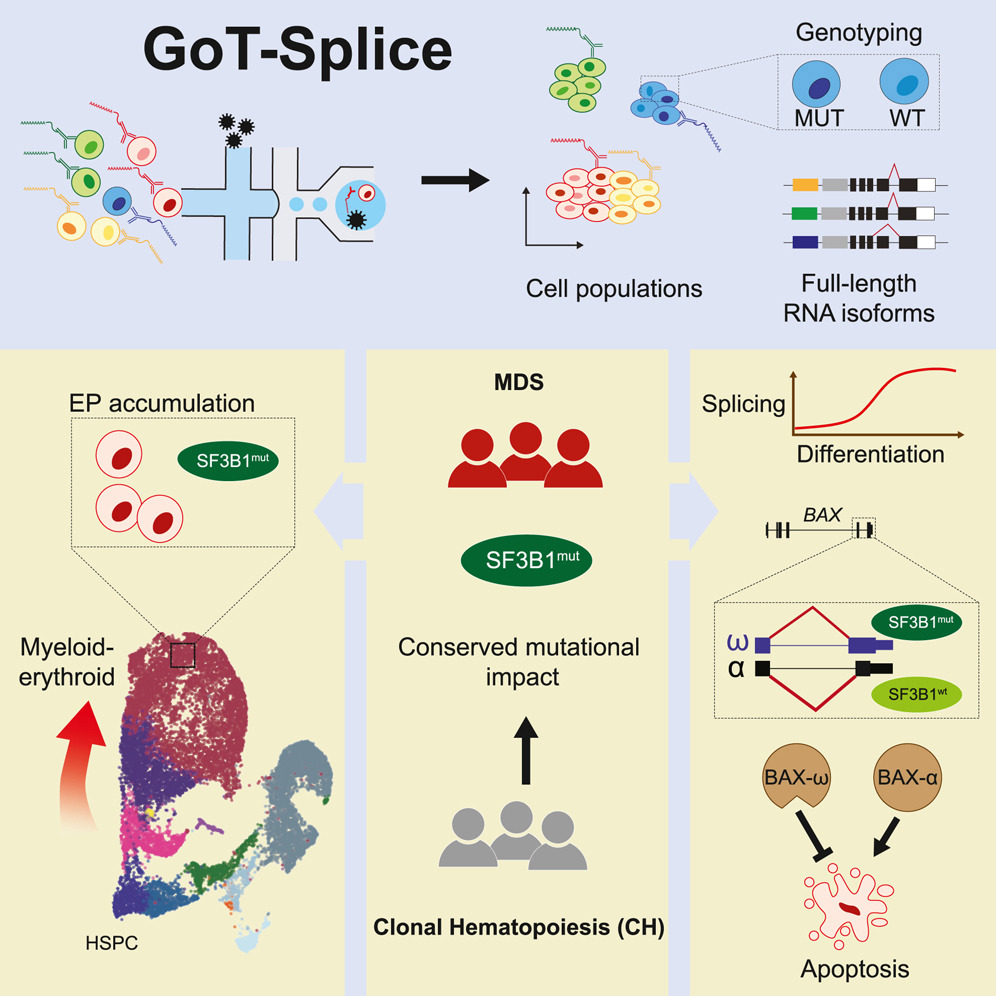
✂️Splice splice baby!✂️ New paper just out in Cell Stem Cell integrating Oxford Nanopore long read single-cell isoform mapping into our broader genotype-phenotype vision for clonal mosaicism ! authors.elsevier.com/a/1hat96tu0Cm5…

Two more openings in our applications development team. Come help us build new computational methods and software to fully leverage the tremendous recent improvements in Oxford Nanopore sequencing! ejnh.fa.em2.oraclecloud.com/hcmUI/Candidat…



Now, Oxford Nanopore only approaches are producing the best t2t automated assemblies ever produced with 30 contigs and 5 scaffolds! 🤯🤯🤯🤯 #nanoporeconf

