Christine He
@christine_y_he
Genomic applications @nanopore 🧬 | bioinformatics, metagenomics, microbiomes 🦠 | previously postdoc @ucberkeley, PhD @georgiatech
ID: 40100162
http://christine-he.webnode.com 14-05-2009 22:13:22
202 Tweet
625 Followers
718 Following

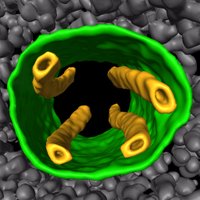
Excited to share our new study with the Müller and SchullerJM labs, online @nature: nature.com/articles/s4158… This is T. kivui, an anaerobic bacterium. It uses hydrogen energy to store #CO2. How does it do it? #CryoET revealed membrane-anchored bundles of the enzyme HDCR 🍩 1/🧵




It's parasites all the way down! John Eppley Elaine Luo Steve Biller et al used Oxford Nanopore sequencing to characterize mobile genetic elements that **hijack phage capsids** to shuttle themselves between hosts in the marine microbial environment.

Have you ever wished to sequence SPECIFIC bacteria but NOT the vast background in #microbiome? Today, we introduce *mEnrich-seq*: #methylation-guided enrichment sequencing of bacterial taxa of interest from microbiome. 4yr project led by Annabel Cao! biorxiv.org/content/10.110…🧵1/10

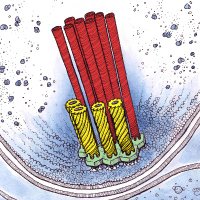
Asgard archaea reignited discussions about the tree of life 🦠 But do they have eukaryotic features? YES!! 🤩 Check out our new paper nature by Florian Wollweber and Jingwei Xu, in an exciting collaboration with the Schleper lab Schleper Lab Vienna. nature.com/articles/s4158… 🧵1/4



Very proud to share that my first-author paper refuting the existence of a human blood microbiome is now out in Nature Microbiology: nature.com/articles/s4156… Huge congratulations to all co-authors and special mention to Niranjan Nagarajan and Minghao Chia. See quoted for a summary🧵.


Looking for a point + click platform for analyzing sequencing data? CZ ID now supports metagenomic analysis with Oxford Nanopore ✳️ Upload Nanopore sequencing data ✳️ No coding required ✳️ Interactive reports ✳️ Download reports Learn more about the pipeline bit.ly/42TCIQX







It hasn't been long since I last quantified Oxford Nanopore accuracy, but the release of Dorado v0.5.0 demanded another test: rrwick.github.io/2023/12/18/ont… Biggest improvement I've seen in a while! Most of these ONT-only bacterial genomes are now >Q60 😲