Justin Cosentino
@cosentiyes
ml+acgt @googleai; cs @tsinghua_uni & @swarthmore
ID: 409681626
http://cosentino.io 11-11-2011 02:38:27
188 Tweet
236 Followers
415 Following



Release of DeepVariant v1.2. Refactor by Maria Nattestad modularizes multi-sample components, improving DeepTrio accuracy. Accuracy improvements for PacBio HiFi, especially for newest PacBio chemistry. Updates to python, TF libraries improves speed. github.com/google/deepvar…


Excited to share new phenotyping methods for COPD, improving COPD GWAS Collaborative work with IU and BWH, co-authors Justin Cosentino, B Besaz , @babak_alipanahi, Z McCaw, last author Farhad Hormozdiari, eng manager C McLean. Paper: medrxiv.org/content/10.110… Code: github.com/Google-Health/…


We’re excited to describe new Google Health work that makes phenotyping more scalable. An unsupervised deep learning method, REpresentation learning for Genetic discovery on Low-dimensional Embeddings (REGLE), to perform GWAS on high-dimensional clinical data.


Excited to share new methods for cardiovascular phenotyping from accessible inputs (ECG and PPG) and scalable unsupervised methods. Co-authors: Yuchen Zhou @cosentiyes, Ted Yun, last author B Besaz & Farhad Hormozdiari, eng manager C McLean. Paper: medrxiv.org/content/10.110…


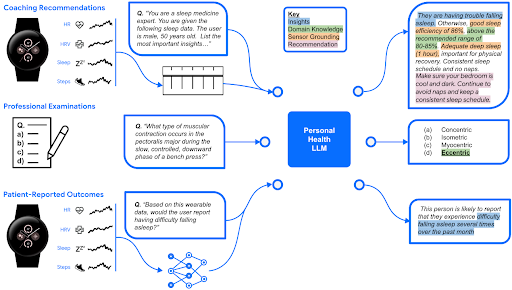
Google’s research on a Personal Health Large Language Model (PH-LLM) is now published in Nature Medicine Magazine!