Yupei You
@youyupei
Postdoctoral researcher at the Walter and Eliza Hall Institute in Matt Ritchie’s lab.
Research topic: Genomics | Transcriptomics | Long-read sequencing | Singl
ID: 1672455198
15-08-2013 07:09:36
34 Tweet
87 Followers
104 Following

Our latest preprint and software "NanoSplicer: Accurate identification of splice junctions using Oxford Nanopore sequencing" led by Yupei You in collaboration with Mike Clark biorxiv.org/content/10.110… github.com/shimlab/NanoSp… Oxford Nanopore 1/3
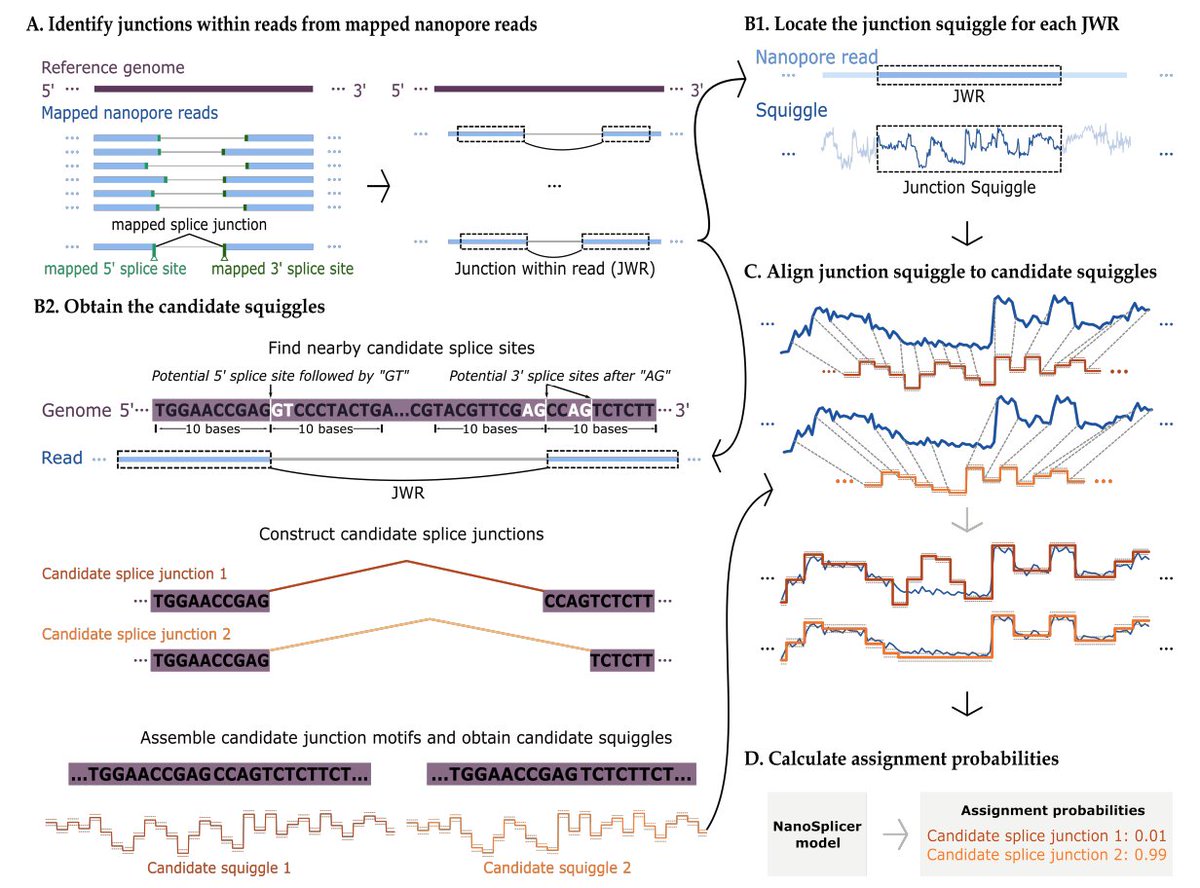


Thanks again to Heejung Shim for giving a virtual seminar on 'NanoSplicer: Accurate identification of splice junctions using Oxford Nanopore sequencing'. If you missed it, you can watch here: youtube.com/watch?v=3YNGnn…

Long-read scRNA-seq is awesome for studying RNA isoforms. But Oxford Nanopore has relied on matched short-reads to identify cell barcodes and assign reads to cells. Pleased to share BLAZE, a new tool to identify cell barcodes from nanopore scRNA-seq. Short-reads not required. 1/4


Amazing week for #DeepLearning in #spatial #singlecell biology, with 2🔥new Graph Neural Networks methods! 1.STELLAR🇺🇸 Jure Leskovec: a cell type annotation & discovery atlas-type framework 2.NCEM🇪🇺 Fabian Theis: an approach to infer cellular communication patterns Deep dive below🧵




Our tool BLAZE for identifying cell barcodes from Oxford Nanopore scRNA-seq is now published in Genome Biology genomebiology.biomedcentral.com/articles/10.11… Lots of improvements from preprint -faster -high sensitivity mode -output to run empty drops to save/discard cells -robust to different read depths




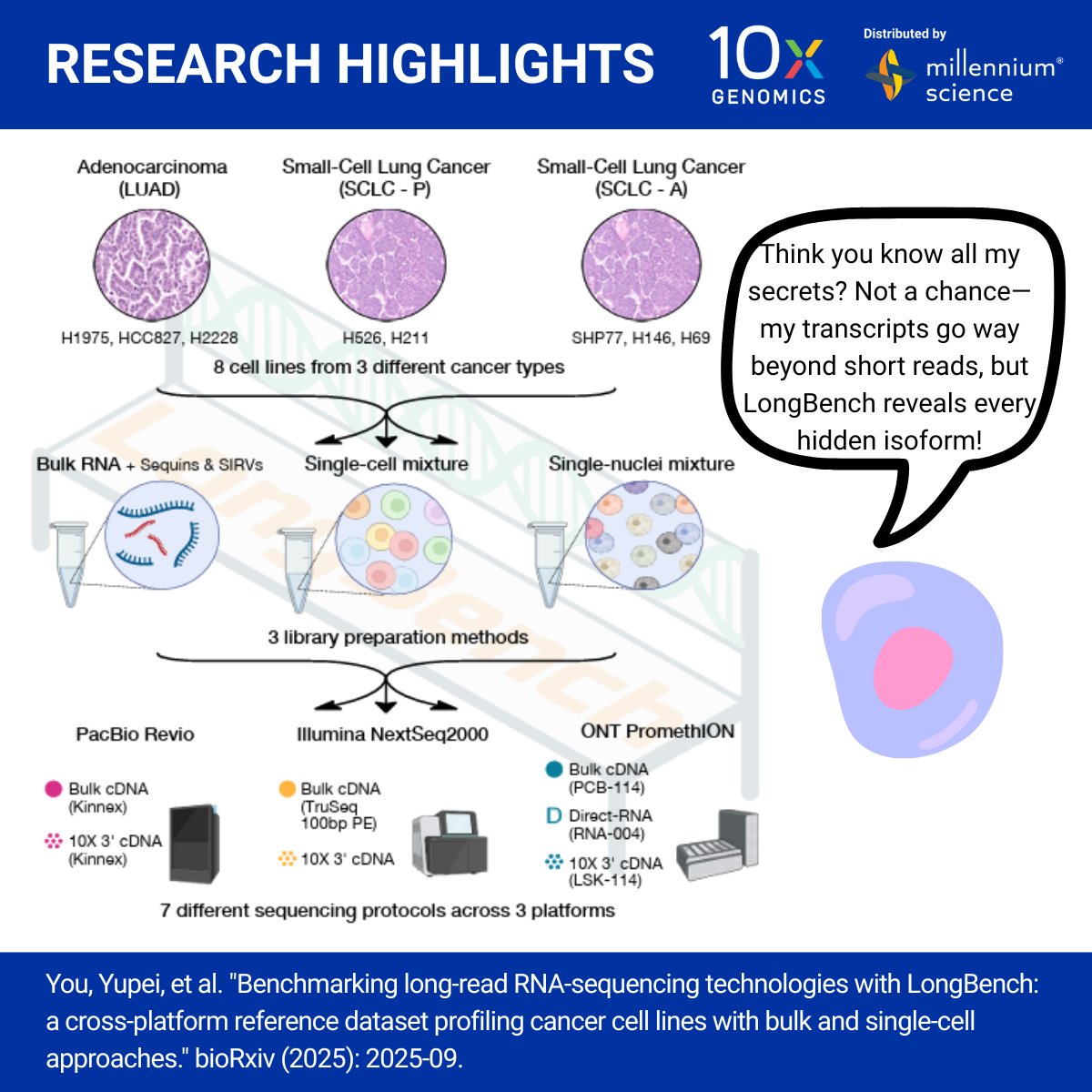
Benchmark long-read analysis with 10x Genomics + LongBench. The LongBench resource from Matt Ritchie's lab, led by Yupei You showcases how 10x Genomics single-cell and single-nucleus workflows reveal full-length transcripts and uncover isoform diversity.