Don't get tied up in pseudoknots!
Project ideas for the coolest computational #RNA #hackathon are due Friday!
Travel grants avail for #RNA 2020 ppl
--> hackseq.com <--
#Bioinformatics The RNA Society Jr RNA Scientists RNA Bioscience Initiative | CU Anschutz Yale Center for RNA Science and Medicine RNAcentral R-Ladies Vancouver BF Francis Ouellette



1)MicroRNA-146a-5p, Neurotropic Viral Infection and Prion Disease (PrD).
2)mRNA Vaccine teaches our cells how to make protein
3/4)Pseudoknots in prion protein mRNAs confirmed by comparative sequence analysis and pattern searching
#coronavirus #clotshot #grapheneoxide #vaccine
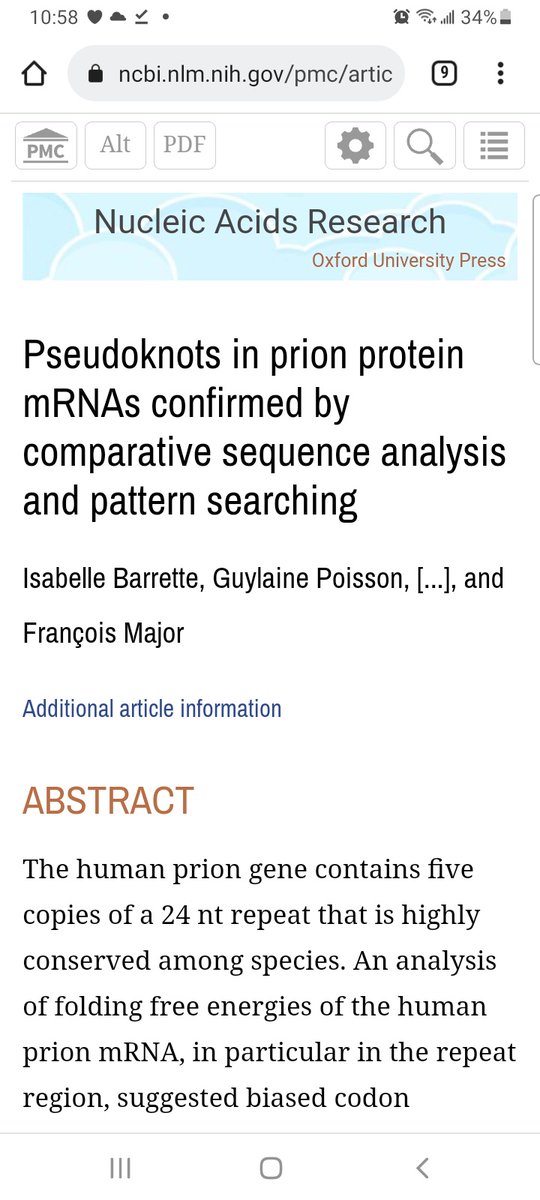


Biotite 0.25.0 provides new tools for structure analysis: Measure nucleic base stacking, create dot-bracket notations, identify pseudoknots, calculate PEOE partial charges and more! Below you can see the charge distribution in a molecule, created with Matplotlib. #bioinformatics


An algebraic language for RNA pseudoknots comparison bmcbioinformatics.biomedcentral.com/articles/10.11… #bmcbioinformatics

Prediction of RNA secondary structure including pseudoknots for long sequences ift.tt/2WAsnvZ #bioinformatics

OK, Markus Deserno, you asked what led me to write a paper on pseudoknots and alpha helices. Why would I, as a membrane biophysicist, be interested in whether a structure misfolds if you start from one end vs. the other? Here is a thread! 1/n


How is the function of #biomolecules impacted by #riboswitches and pseudoknots? Our Mathematical Biocomplexity division studies the topological complexity of interacting systems in the hopes of gaining insight into #genes with complicated structures. ow.ly/bBcZ50D0Ajk




Chemical probing data for 4600 Eterna player-designed #RNA sequences is now publicly available in the RNA mapping database rmdb.stanford.edu for #datascience #ML needing a large dataset of #pseudoknots to train 3D prediction algorithms.



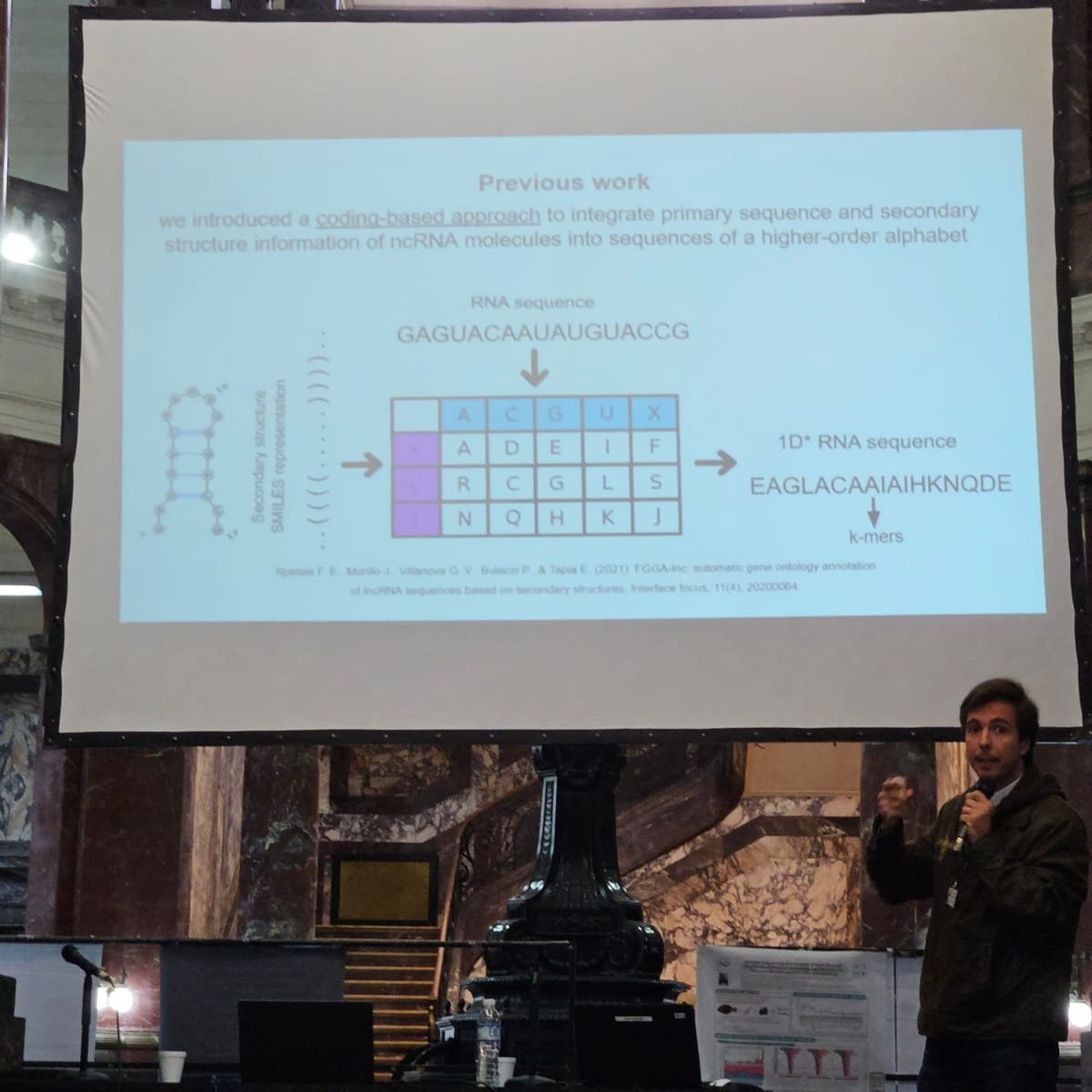
Ignacio Labari from Argentina 🇦🇷 and a member of #RiaBio is bringing us a fascinating session on Day 2 of the conference: 'A music-inspired codification of ncRNAs with pseudoknots.' 🎶🧬📚 #Bioinformatics #RiaBio 2023 #ScienceInnovation



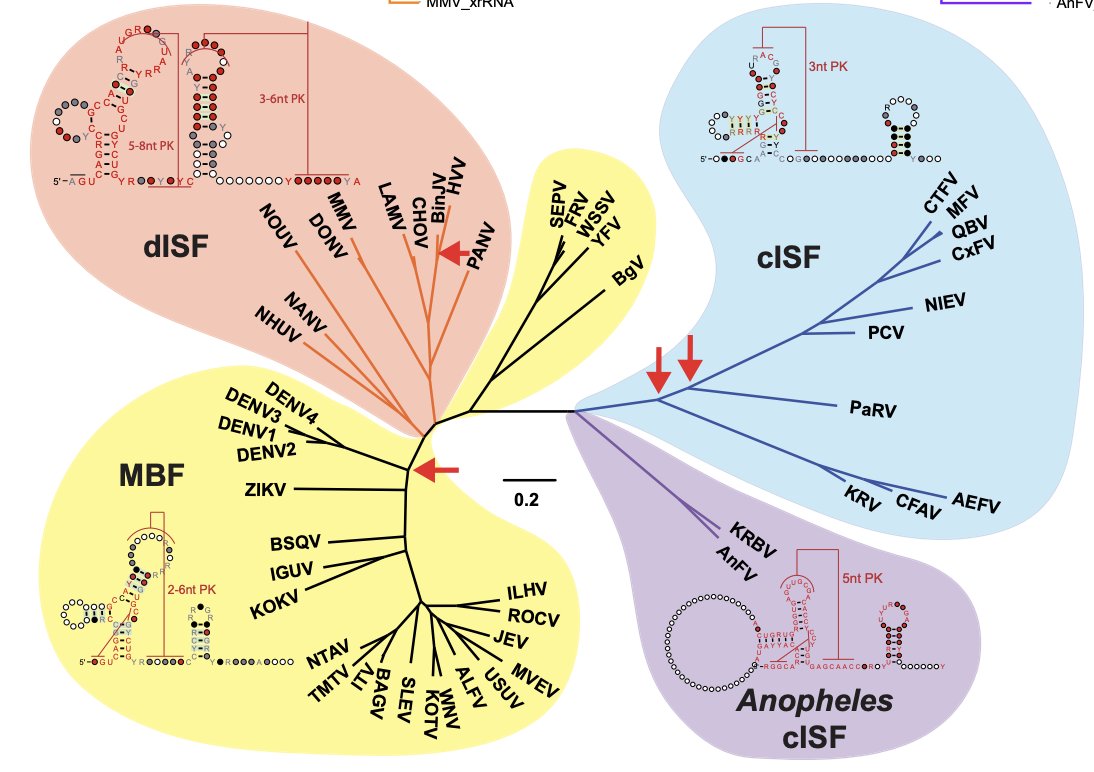
Work from Dr Andrii Slonchak, Alex Khromykh lab published in Nature Communications examines structural features of 3’UTRs in classical insect & dualhost #flaviviruses (cISF/dISF). Elaborate SHAPE data show c/dISF have multiple xrRNAs and dISF contain novel pseudoknots
nature.com/articles/s4146…