Gerald Schwank
@schwanklab
Associate Professor @UZH_en.
We develop genome editing approaches for personalised treatment of rare diseases and cancer.
ID:943028500570746880
https://schwanklab.org/ 19-12-2017 08:01:29
104 Tweets
948 Followers
160 Following

I'm excited to talk @ the #AMLDEPFL2024 conference today together w/ Nicolas Mathis. Join us and Amina Mollaysa Charlotte Bunne Bruno Correia Manuel S. for AI in Genomics in Garden4C from 11 am on. You'll also find us at the AI x bio after-party lu.ma/amldepfl 🧬x🤖


Coming soon! Latsis Symposium on Genome Engineering #Zurich #switzerland June 13-14. latsis-genome-editing.ethz.ch Great lineup Treutlein lab Ben Kleinstiver Raffaella Di Micco Sek Kathiresan MD Janice Chen Charles Gersbach Cejka Lab Teresa Davoli Jolanda van Leeuwen Cereseto Lab Anna Obenauf Prisca Liberali

Great news today for IHB. I warmly welcome Juergen Knoblich as new director and head of the institute. Thrilled to see the next step of our strategic evolution come to fruition and our ambitions to expand into basic biology realised. See you soon in Basel The Knoblich Lab!

If you want to do Synthetic Biology and you love lakes and mountains you have to apply! UZH’s Faculty of Medicine seeks candidates for a Professorship in Synthetic Biology/Protein Engineering (open rank). tinyurl.com/9bka23pb #AcademicJobs #SyntheticBiology #UZH




Impossible to do the field justice on a few pages but we (Jonathan Gootenberg Omar Abudayyeh Jeremy Walker Julia Joung Luke Koblan) tried our best to give an overview of current CRISPR technologies in Nature Reviews Molecular Cell Biology rdcu.be/dxDD9


Just shared at Keystone Symposia a new @ArcInstitute discovery of the bridge RNA recombinase mechanism: a new class of natural RNA-guided systems that retains the key property of programmability from RNAi and CRISPR while enabling large-scale genome design beyond RNA and DNA cuts


exciting new work from Matt Durrant #hiring and Nicholas Perry & co at Arc Institute!
bridge RNAs direct modular and programmable recombination of target and donor DNA
🧬x🧬
if you’re into transposons, CRISPR, MGEs, recombinases, or programmable biology, you’ll want to check this out:

Extremely excited to kickstart my lab TU München with Emmy Noether support from DFG public | @[email protected]. I’m very grateful to J. Keith Joung, my friends in Boston, and to A. Moretti, K.L. Laugwitz & C. Kupatt in Munich.
More details & website coming soon. #CRISPR 🫀


🚨New preprint🚨Today, we're excited to share our latest study, 'Predicting prime editing efficiency across diverse edit types and chromatin contexts with machine learning' via bioRxiv! Gerald Schwank Krauthammer Lab Bas van Steensel lab UZH Science
biorxiv.org/content/10.110…
🧵

In our recent preprint Nicolas Mathis and collaborators developed new computational models predicting the impact of pegRNA design and local chromatin context on prime editing. Access the tools at pridict.it. Dive into the thread or read our preprint for more details:

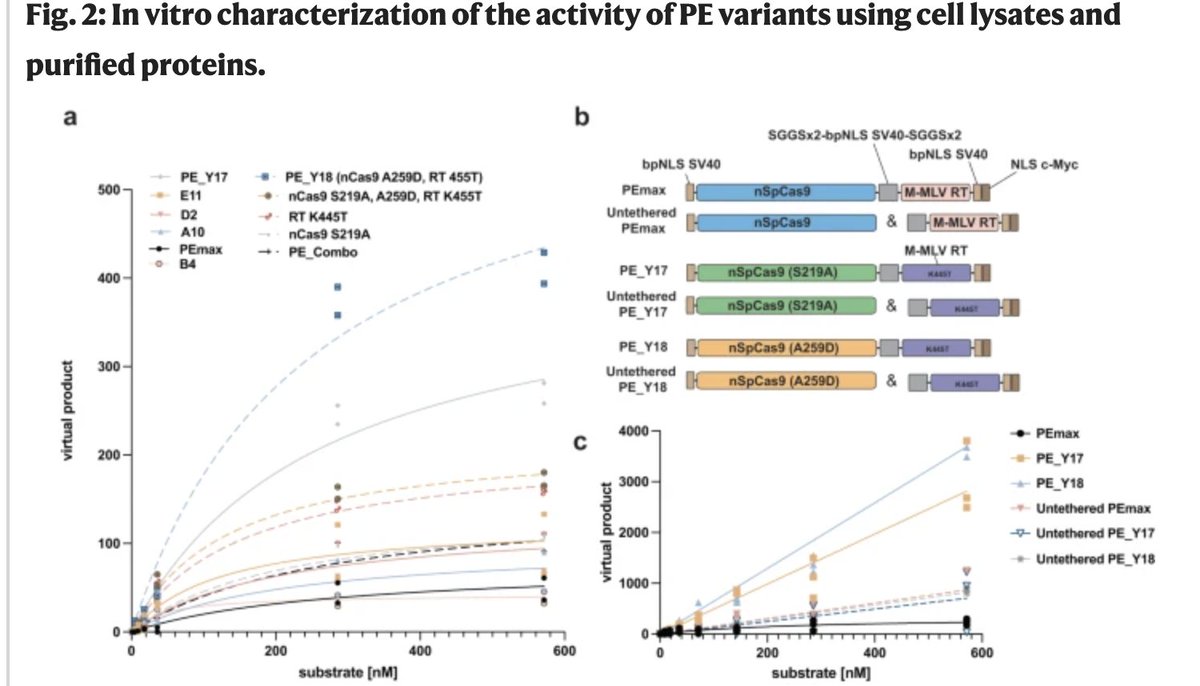
evoCjCas9 - a new addition to the genome editing toolbox! Big congrats to Lukas Schmidheini and everyone from the Gerald Schwank and Martin Jinek lab supporting this project.


Just out Science Magazine
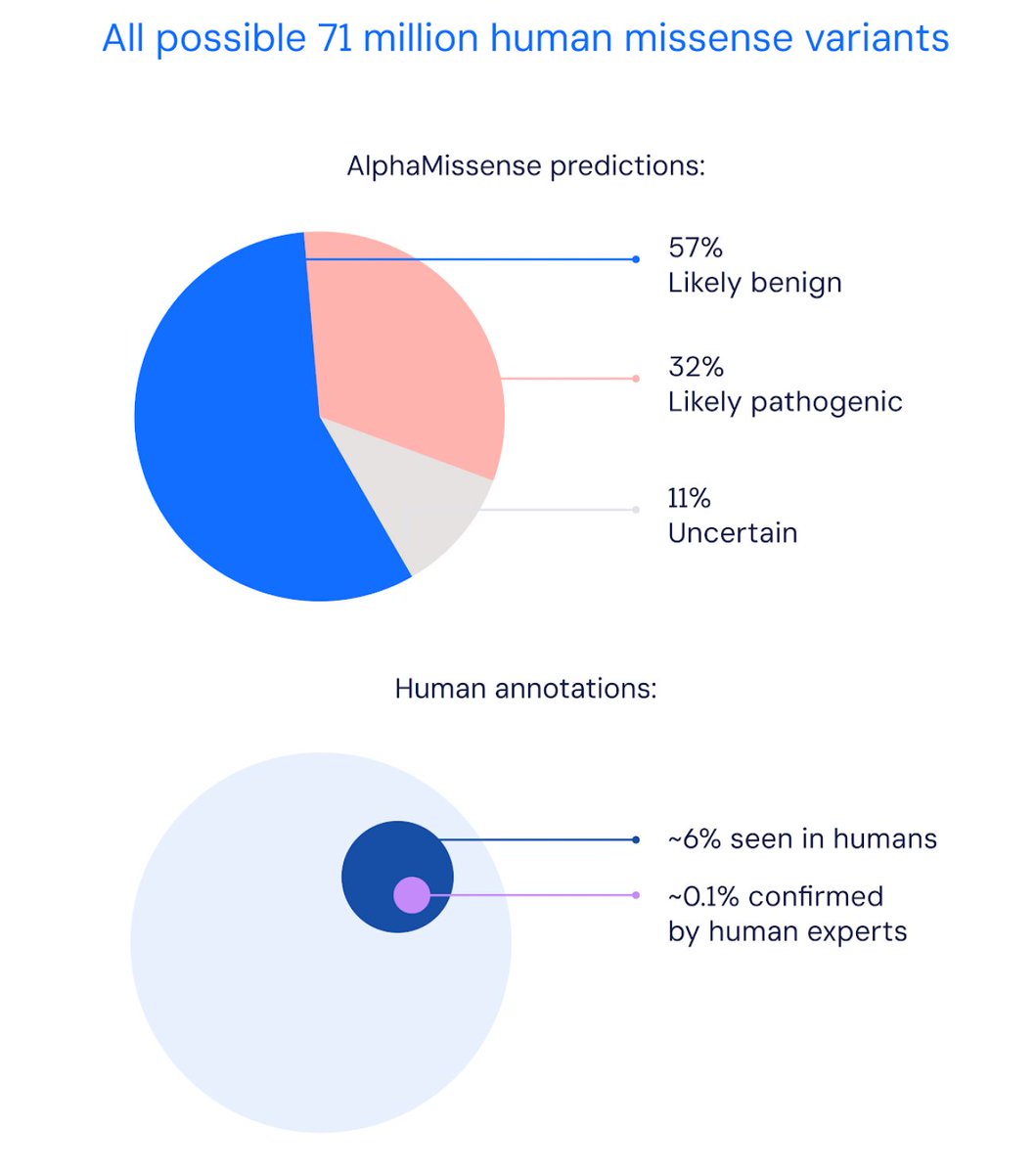
#AlphaMissense —building on #AlphaFold —based on unsupervised learning #AI , predicts impact of all 71 million human missense variants for disease-causing potential, across entire human proteome; open-source👍
science.org/doi/10.1126/sc… @jun90cheng Žiga Avsec