Herczeg Robert
@robertherczeg
Data scientist, PhD in ecology. I R :)
ID: 3300246153
http://www.rdata.hu 27-05-2015 09:59:22
125 Tweet
81 Followers
326 Following



Just got the The Carpentries (at @[email protected]) instructor certificate. We will likely organize an official #datacarpentry and/or #softwercarpentry workshop during autumn. Cannot wait :)

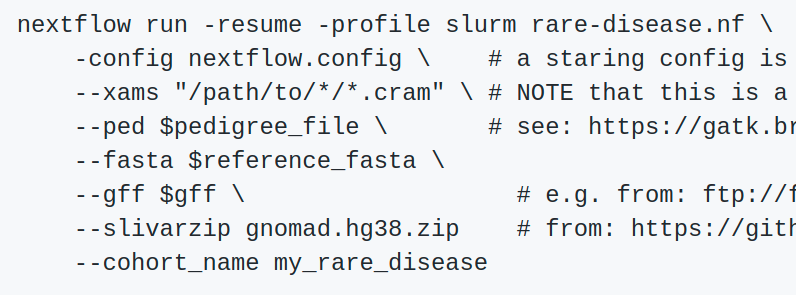
Luca Cozzuto from the Centre for Genomic Regulation (CRG) Bioinformatics Core @ CRG Describes a Nextflow pipeline for processing nanopore sequence data Challenges exist - how to remove very large intermediate files - how to use GPUs in the best way (will probably use cloud for rather than cluster) #nfcamp19



2020 International #Bioinformatics #conference in #Thessaloniki, Greece. Mark your calendars for November 20-22, 2020. More to come soon! #Hbioinfo2020 Elixir Greece ELIXIR Europe ISCB Student Council #ISCB


Thank you to GISAID Initiative and the Bioinformatics Research Group at Szentagothai Research Centre, we now have 2 sequences from Hungary at nextstrain.org/ncov?f_country…. This new sequence nests separately from the first, grouping with viruses from the Netherlands and Switzerland.


Thanks to #opendata sharing on GISAID Initiative, nextstrain.org/ncov is updated with 59 new #COVID19 #SARSCoV2 #hCoV19 sequences! These are: - USA (19: 2 WI, 17 CA) - Brazil (1) - Hungary (1) - Spain (1) - France (23) - First 2 seqs from Algeria! - First 12 seqs from Senegal! 1/4



The second session in the Galaxy Project-ELIXIR webinar series will present the analysis of the #SARSCoV2 genome: is.gd/rYsll4 Join us on Thursday 7 May at 17.00 CEST, ❗️Register: is.gd/UzCMW9 #ELIXIRvsCOVID19 de.NBI / ELIXIR-DE Galaxy Australia ELIXIR Belgium


Historical moment in #filovirus research, sequencing the complete genome of #lloviuvirus in 50 minutes after a decade. Oxford Nanopore Tóth Gábor Endre #filoviridae #emergingdisease #bat #virology


I have the opportunity to learn within the #HelloAIRIS training by EIT Health from 15 June to 10 August 2020 we'll learn about #AI in #Healthcare. Looking forward to hearing from esteemed experts from Leitat, GE HealthCare, and KTH Royal Institute of Technology