
Olli Dufva
@ollidufva
MD, PhD, postdoctoral fellow @sangerinstitute, previously @helsinkiuni. Functional genomics in immunology and cancer
ID: 813802186249555969
27-12-2016 17:42:15
1,1K Tweet
1,1K Followers
848 Following

So excited to share this work! It has been incredible to work with Tom Thomas Norman and the Norman Lab members to put this together. We made several key advances, did truly massive experiments, and learned about fibroblast biology. Big shout out co-author Rico Chandra Ardy!

🚨𝐇𝐞 𝐋𝐚𝐛 job opportunity UC San Francisco 🚨: I'm hiring TWO postdocs - an experimentalist and a computational biologist. Join us to use cutting-edge single-cell and spatial technologies to unravel the mysteries of human tissues. opportunities.ucsf.edu/content/open-p…
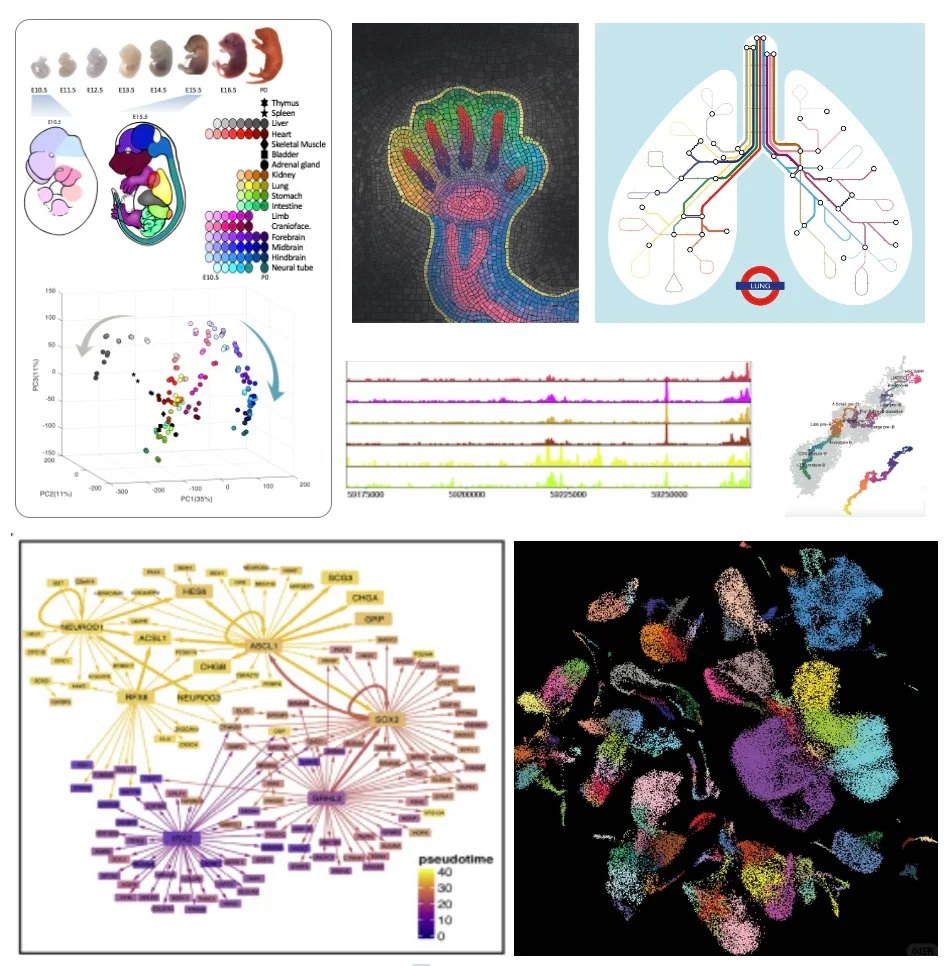

It was great to work with Nature Methods on this piece about foundations models for cell biology, while we are not there yet, the future is bright. Effective training of these models guided by tasks and evaluations that are non trivial with careful benchmarking against existing





Our new #paper is out in Nature Immunology. Applying #spatial transcriptomics at subcellular resolution to 130 ovarian #cancer tumors, followed by data-driven experimental design and high content #CRISPR screens to map immune evasion in ovarian cancer nature.com/articles/s4159… (1/15)

Major effort from the lab (💪Robert Frömel), finally on biorxiv: We built 60,000 fully synthetic enhancers and measured their cell state specific activity during blood stem cell differentiation. Why did we do that and what did we learn🧵? biorxiv.org/content/10.110…

Made it to the other side! Had an absolutely fantastic PhD defence day, with great examination by Prof. Eelco de Koning and celebration with friends and colleagues


Friends! I am super-excited to share our story on KLHL6 in this @BCD_aacr 📣KLHL6 targets the B-cell receptor and lymphoma mutations perturb this homeostasis! 🔓Read doi.org/10.1158/2643-3… 🪩Joint editorial in Cancer Discovery doi.org/10.1158/2159-8… 🧵Let’s go


Excited to publish this by the NCI/Cambridge team Our approach building from KLHL6 up and their pola-V CRISPR-screens moving down from CD79B come to the same conclusion for KLHL6 role doi.org/10.1158/2159-8… Congratulations Sean Corcoran Daniel Hodson and the team!


🌟 I am thrilled to share that our perspective article on Molecular Connectomics, co-authored with Nadav Yayon, Kerstin Meyer, and Sarah Teichmann Teichlab FMedSci FRS, has been accepted for publication in PLOS Biology. 1/n

One of the last stories from my postdoc work with Howard Chang and William J. Greenleaf is out today Science Magazine led by Laksshman Sundaram. scATAC-seq of 8 different tumor types from TCGA. science.org/doi/10.1126/sc…



Commentary on our recently published T-ALL cancer genomics research in Nature Genetics by Safia Danovi nature.com/articles/s4158…

Please have a look at our g2g framework on aligning cell trajectories developed by Dinithi Sumanaweera, Chenqu Suo Ana-Maria Cujba and others in the team! Congrats! Wellcome Sanger Institute Cambridge Stem Cell Institute Cambridge University


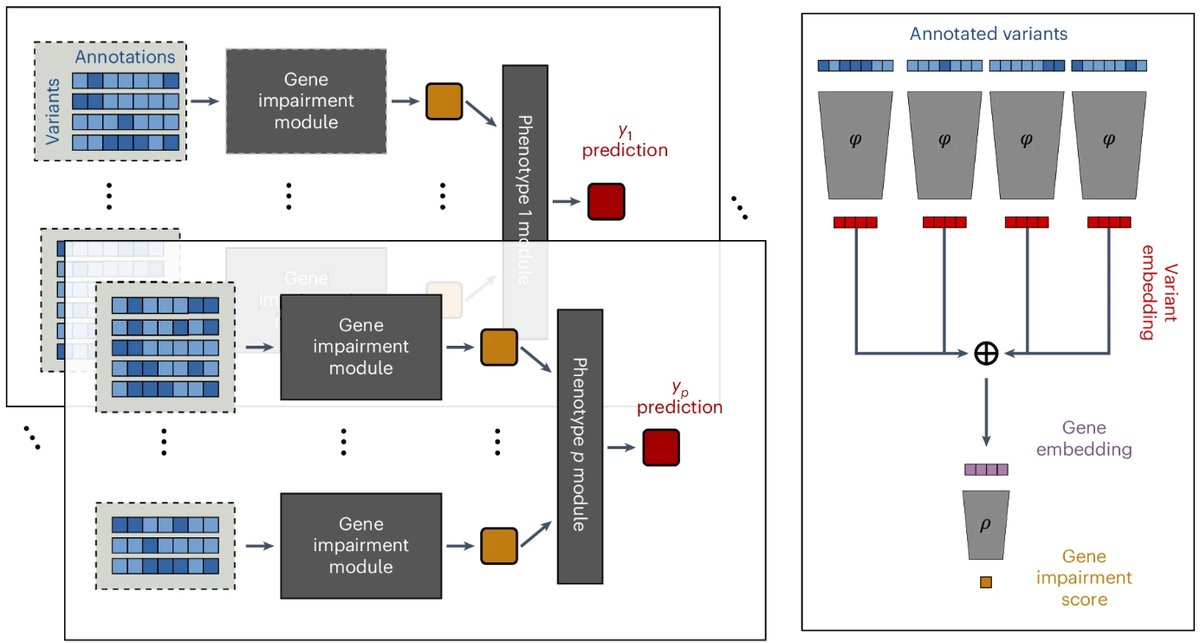
🧵 1/9 Delighted to announce that #DeepRVAT is out today in Nature Genetics! Here's a recap of our tweetorial from the preprint, with updates for some exciting new results. It's been a joy to work with Eva Holtkamp, as well as gagneurlab and Oliver Stegle.

Tremendously proud to share my fellowship work, online today in Nature Medicine. We tried to understand why CAR T therapy for AML doesn't work as well as we hoped it would, and how we can solve that. 👇🧵1/10 Penn Medicine CHOP Research Saar Gill nature.com/articles/s4159…