Fredrik Dyrkell
@lexicallyscoped
Programming languages and bioinformatics enthusiast.
CTO at 1928diagnostics
ID: 992593801
http://www.lexicallyscoped.com 06-12-2012 07:36:53
7,7K Tweet
489 Takipçi
336 Takip Edilen


Fantastic collaboration with Kiki Xanthopoulou Sebastian Alexander Fuchs Alexander Dilthey pmrCAB Recombination Events among Colistin-Susceptible and -Resistant Acinetobacter baumannii Clinical Isolates Belonging to International Clone 7 | mSphere journals.asm.org/doi/10.1128/ms…
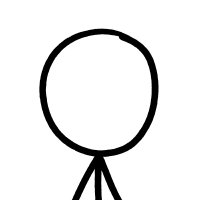








New paper led by @mbhall88 and collaborators! Evaluating Oxford Nanopore as a replacement/complement for Illumina for TB: drug resistance and clustering. Spoiler: you can get equivalent clustering and identical resistant predictions (with 30x depth). 1/n medrxiv.org/content/10.110…

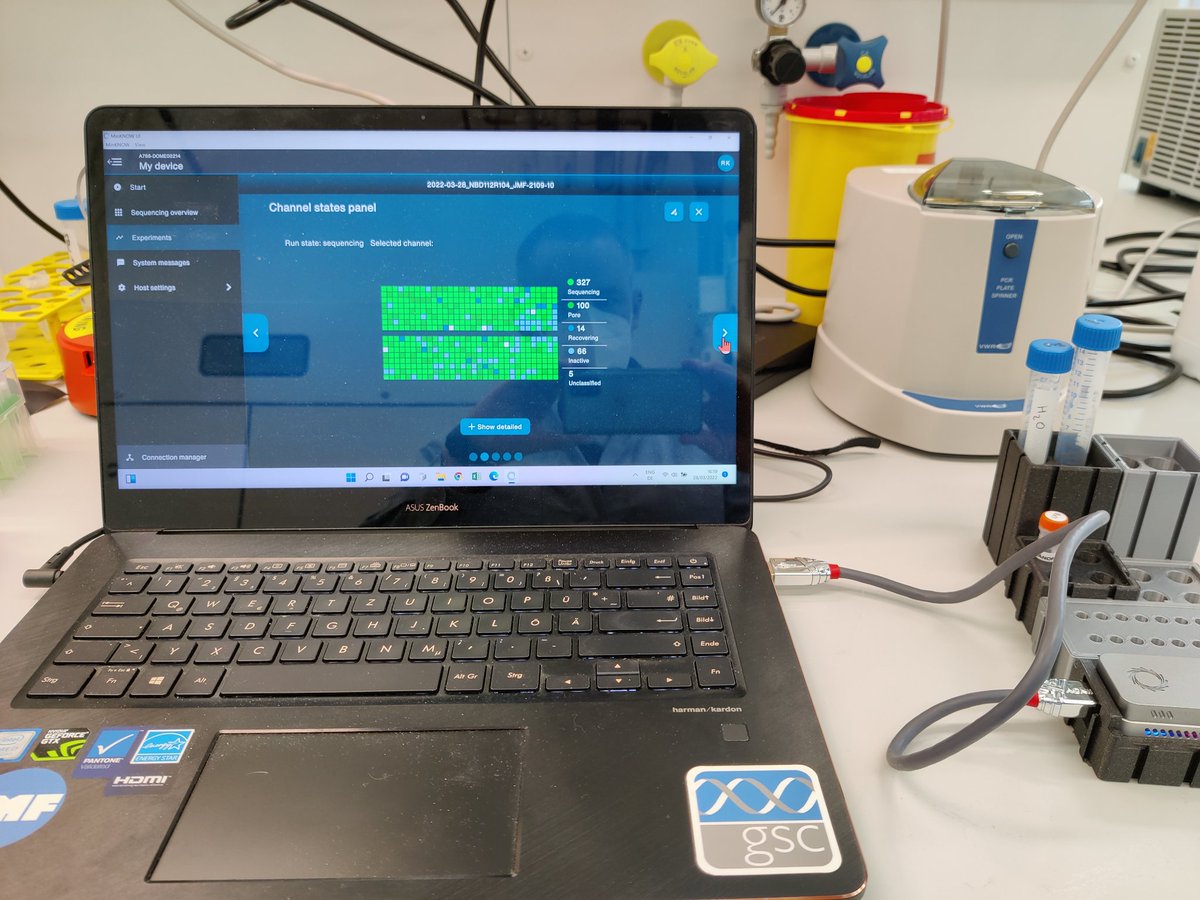
Picking read size and running adaptive sampling directly in Oxford Nanopore minknow is now available 🎉😁 #SQK-NBD112-24 prep of pure cultures. Looking forward to a RBK version for faster prep.


@Johngorzynski, Kishwar and I are excited to present the details of our ultra-rapid Oxford Nanopore whole genome sequencing pipeline (nejm.org/doi/full/10.10……) published in Nature Biotechnology nature.com/articles/s4158… today! The pipeline produces a genetic diagnosis in under 8 hrs.




Hey Dave Grossman isn't here, Guybrush invited us to meet him at the Scumm Bar this evening. You coming? I'm going to show up fashionably late. #MonkeyIslandMonday #ReturnToMonkeyIsland

Amazing story about a rubik cube themed jigsaw puzzle and a solver written in Zig, with a dash of raylib technologies for visual feedback. zig.news/shadeops/zig-a…

Presenting sylph (github.com/bluenote-1577/…), a fast, precise metagenomic profiler. Work done with Yun William Yu. sylph is highly accurate at the species level and takes ~1 minute and 16 GB RAM to profile against 85k prokaryotes and 2.9 million viral genomes. 1/10


Control Structures, English translation of Collège de France lectures by Xavier Leroy discuss.ocaml.org/t/control-stru…