M. Julius Hossain
@juliushossain
Head of Image and Data Analysis Group, Centre for Cancer Immunology, University of Southampton
ID: 415988599
https://www.southampton.ac.uk/people/5zcf55/doctor-m-julius-hossain 19-11-2011 02:48:21
33 Tweet
39 Followers
33 Following










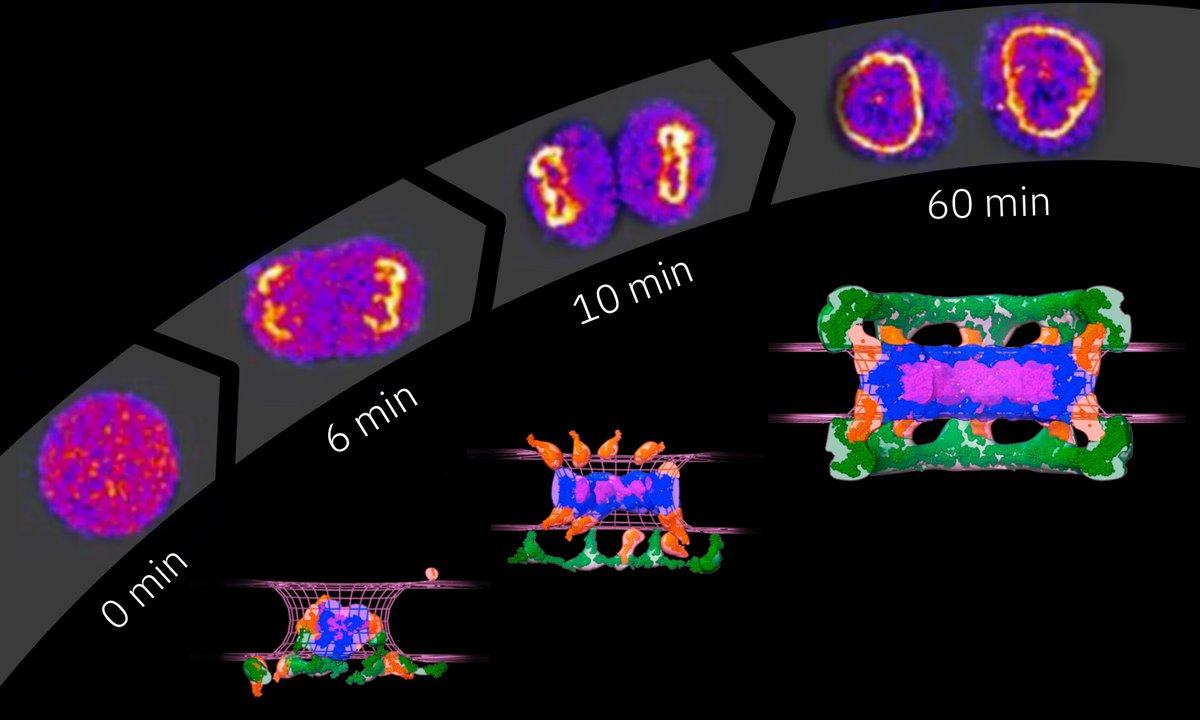
with ClaudiaCattoglio Tjian + Darzacq Group @EllenbergLab Nike Walther Xavier Darzacq Thanks to Nike Walther and @EllenbergLab for the collaboration and for cross-validating the estimates using calibrated FCS-imaging. Numbers in image below. We would appreciate any comments or suggestions.










We are currently welcoming submissions of original research to the “Advances in fluorescence-based medical imaging” Collection in BMC Medical Imaging biomedcentral.com/collections/af… Dr. Inavalli


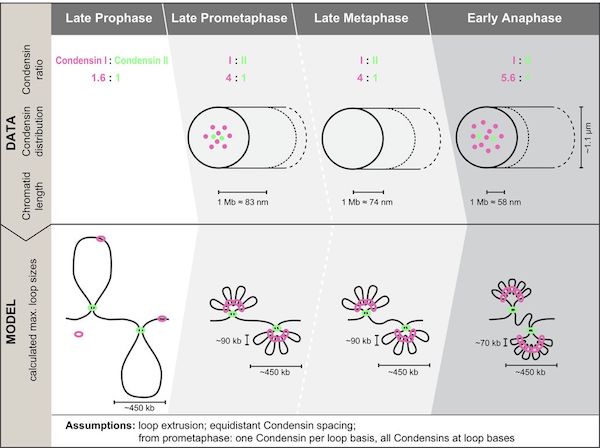
Walther et al systematically fluorescently tag endogenous Condensin subunits and map their abundance, physical spacing, and mitotic dynamics, proposing a three-step hierarchical loop model of mitotic chromosome compaction @EllenbergLab M. Julius Hossain jcb.rupress.org/content/early/…