Melanie Weilert
@melanieweilert
bioinformatics, deep learning, genomics, @ZeitlingerLab ((all views are mine alone))
ID: 1070355717163491328
05-12-2018 16:34:24
157 Tweet
157 Followers
234 Following




Please join us in congratulating Khyati Dalal (Khyati Dalal), @KUMedcenter #graduatestudent in the Julia Zeitlinger (@[email protected]) lab, on a successful #thesisdefense! 👏 Well done.


There is so much talk lately about #ArtificialIntelligence (#AI) – but what exactly is it? Melanie Weilert, Bioinformaticist in the ZeitlingerLab, explains what it is & how it's used at the Institute & in medical #research. #WorldAIWeek MORE: bit.ly/45zKorD


Gorgeous paper, Surag Nair ! Fascinated by the nature of low-affinity OctSox enrichment across transient peaks during reprogramming. Very useful work! biorxiv.org/content/10.110…



The sequence syntax that boosts TF binding also transfers to their signaling TF partners. Excellent work, Khyati Dalal !


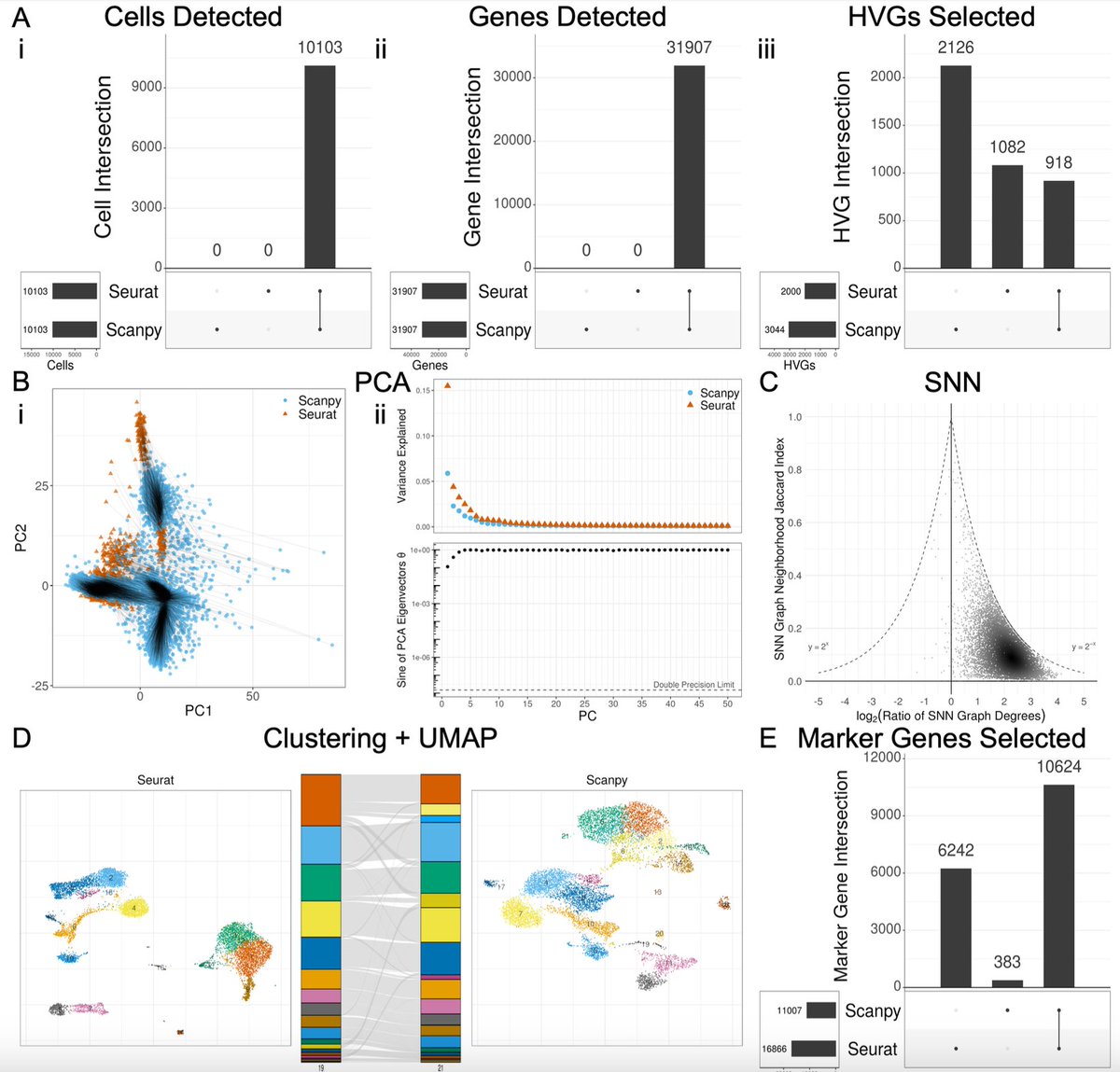
The choice of whether to use Seurat or Scanpy for single-cell RNA-seq analysis typically comes down to a preference of R vs. Python. But do they produce the same results? In biorxiv.org/content/10.110… w/ Joseph Rich et al. we take a close look. The results are 👀 1/🧵



Check out ProCapNet by Kelly Cochran, a robust, local sequence context deep learning model of base-resolution transcription initiation profiles (PRO-cap) that provides deep insights into the cis-regulatory code of initiation & more ... biorxiv.org/content/10.110… 1/

Last week we kicked off our first #OpenMicScience of the summer with presentations from Amanda Lawlor, A.J. Treichel, and Melanie Weilert. 👏 #science #biology


Excellent talk by Michael Hoffman @michaelhoffman.bsky.social predicting transcription factor binding. Old school motif-based approaches are not the answer, most TF binding sites lack TF’s sequence motifs. #ISMB2024 1/


It was an honor to defend my PhD thesis today Stowers Institute. I am so appreciative of my advisor Julia Zeitlinger (@[email protected]) for her guidance and mentorship, as well as my committee members Matt Gibson, BazziniLab , Jerry Workman, and Chris Rushlow for their support! What a terrific day!


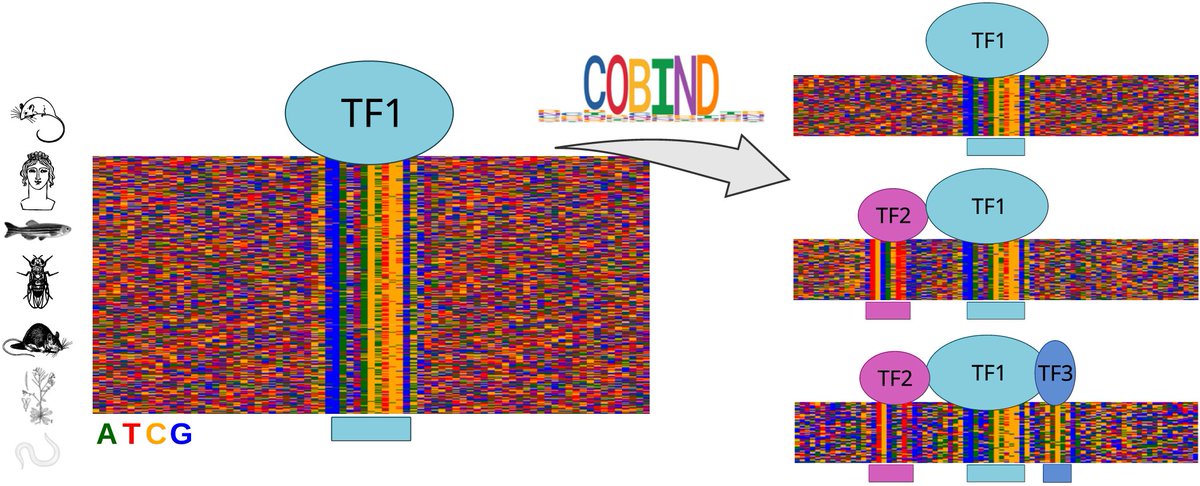
🎉 Excited to share the peer-reviewed version of our manuscript describing COBIND, a tool for revealing TF co-binding patterns using non-negative matrix factorization. Great work led by Ieva Rauluševičiūtė NCMM Nordic EMBL P'ship UiO - Det medisinske fakultet Universitetet i Oslo academic.oup.com/nar/advance-ar… Nucleic Acids Res