
Christian Gnann
@christiangnann
ID: 1094384161291350016
09-02-2019 23:54:51
21 Tweet
12 Followers
32 Following

Wonderful day! After 3 years, probably one of our largest contribution to science is out. #DeepLearning #proteomics #singlecell Matthias Mann Lab Emma Lundberg Andreas Mund BIOMAG

Imaging+LMD+AI+MassSpec = Deep Visual Proteomics ❤️ The possibilities are endless when using imaging to drive supervised or unbiased single cell collection for deep proteomics. Fun project with an amazing team 🤩 Matthias Mann Lab Peter Horvath Towards next gen #pathology!


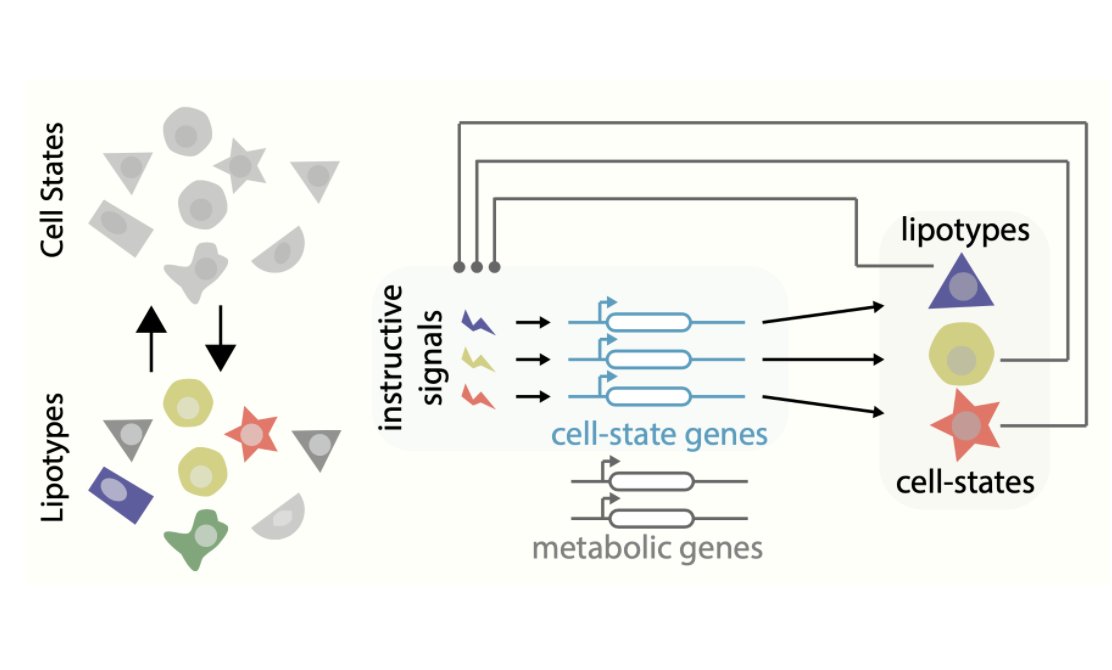
Did you know #lipids control cell identity? Yes, they do! Happy to share our first preprint on #singlecell #lipidomics in collaboration with @GiO_DAngelo and a fabulous team led by Laura Capolupo and Irina Khven. biorxiv.org/content/10.110…


Very proud to present our work "Spatiotemporal dissection of the cell cycle with single-cell proteogenomics” in @nature today. A great (and long) team effort led by Diana Anthony Cesnik, w Human Protein Atlas. A thread... #cellatlas #cellcycle go.nature.com/3sq8sKE


Endogenous tagging + 3D live-cell imaging + mass-spec interactome = opencell.czbiohub.org, an open-source map of the human proteome. So excited for our first release, wonderful collaboration with Matthias Mann Lab Loïc A. Royer 💻🔬⚗️ & many other colleagues. OpenCellCZB


Really excited to share our work that's now online nature, introducing a novel AI approach to integrate confocal images in the Human Protein Atlas (Emma Lundberg) with AP-MS data in BioPlex (Ed Huttlin) to create a unified map of human cell architecture. nature.com/articles/s4158…

Very proud to share our work in nature on how to build a multi-scale hierarchical cell model! With AI we integrate image and PPI data to model the protein communities underpinning cell function. Great collab w @Yue_Qin_UCSD Trey Ideker Ed Huttlin Gygi Lab go.nature.com/30W93uU

Our paper is out today in Science Magazine, I’m stoked OpenCellCZB ! Check out opencell.czbiohub.org: CRISPR tagging + 3D live imaging + IP mass spec to map the human proteome. science.org/doi/10.1126/sc… 👇 tweetorial from our 2021 pre-print.


Deep Visual Proteomics - leveraging imaging and mass spec for deep #spatialomics in native FFPE tissue, in Nature Biotechnology today rdcu.be/cNU70. Great collaboration led by the dream team Matthias Mann Lab Peter Horvath and Andreas Mund -> Towards #digitalmolecularpathology 🌟

I am honored to receive the 2022 Nikon Young Scientist award by DGZ, @DGZ.bsky.social for the research conducted during my PhD time at the Dagmar Wachten & lab lab. Thank you to all the people who have supported me over my journey in #cilia and #flagella science!

Our PhD student @franziska_hibr AnkarklevLab is one of three young researchers taking part in a conversation on #genetics and #genetechnology Nobel Prize Museum on 18 November.

I am happy to share our #bioRxiv preprint where we describe transcriptional heterogeneity of host-pathogen interactions in #Toxoplasma gondii infected immune cells (BMDCs), a great team effort by AnkarklevLab, Barragan Lab and Johan Henriksson! biorxiv.org/content/10.110…

