Andrea Pasquadibisceglie @andpdb.bsky.social
@andpdb
Staff Scientist @Tigem_Telethon
ID: 855737346536468480
https://www.linkedin.com/in/andrea-pasquadibisceglie/ 22-04-2017 10:57:37
754 Tweet
299 Followers
946 Following


So proud to see our work on machine learning + enzyme design just published! doi.org/10.1016/j.cels… Fun collaboration between X, The Moonshot Factory Google DeepMind & Triplebar that we hope can be a template for integrating ML and high throughput screening in protein engineering





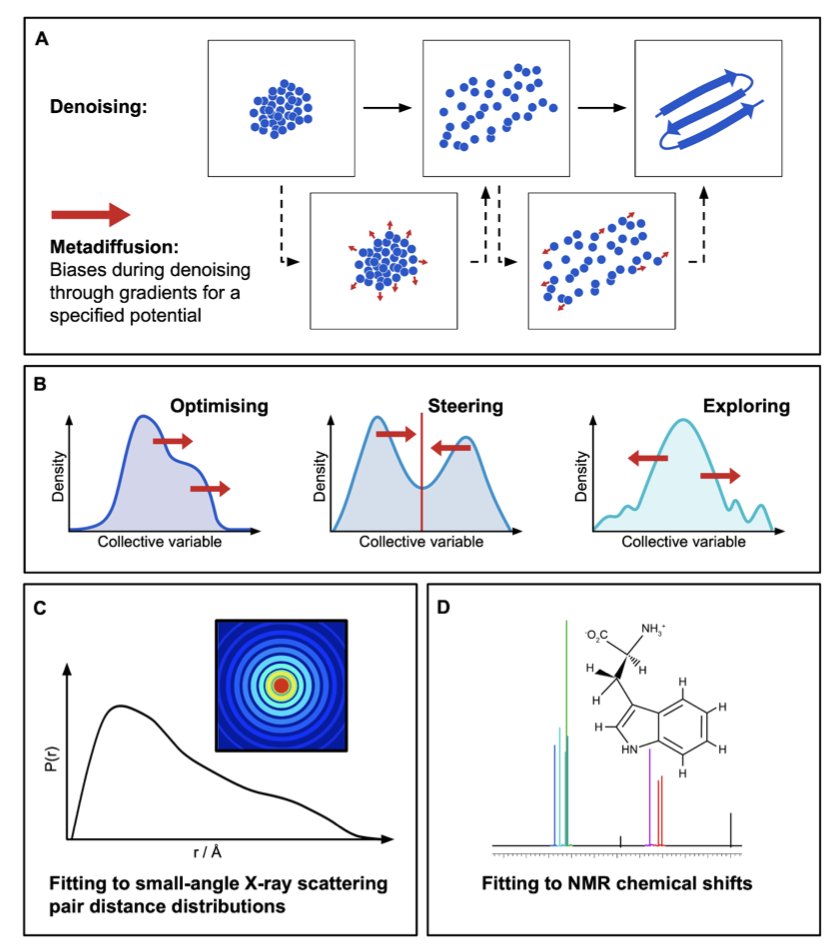
The follow-up work by Enrico Trizio and Peilin Kang is out on Nature Computational Science!🧨 A semi-automatic method to extensively sample "everywhere and everything" (metastable and transition states) "all at once" (in a single simulation) 🔥 📑: nature.com/articles/s4358… Short🧵 IIT






Priyadarshan Kinatukara The MSA is essentially the experimental data used as input to AlphaFold. AF didn't solve the protein folding problem, it solved the graph extraction (from MSA) and 3D embedding problem. Knowing the MSA, lets one analyze if low confidence prediction is simply due to lack of data.