Andreas Krämer
@__kraemer__
Researcher in machine learning and molecular dynamics. @DEShawResearch. Previously PostDoc at FU Berlin and NHLBI (NIH).
ID: 1081625219863851008
05-01-2019 18:55:22
46 Tweet
378 Followers
256 Following

Great new work by Frank Noe Steven Brunton Andreas Mardt BH Tim Hempel et al Freie Universität Mechanical Engineering at UW UW Engineering CPBF - '#Deeptime: a #Python library for #machinelearning dynamical models from #timeseries...' - bit.ly/3rPRHvn #fluids #compchem #dynamics
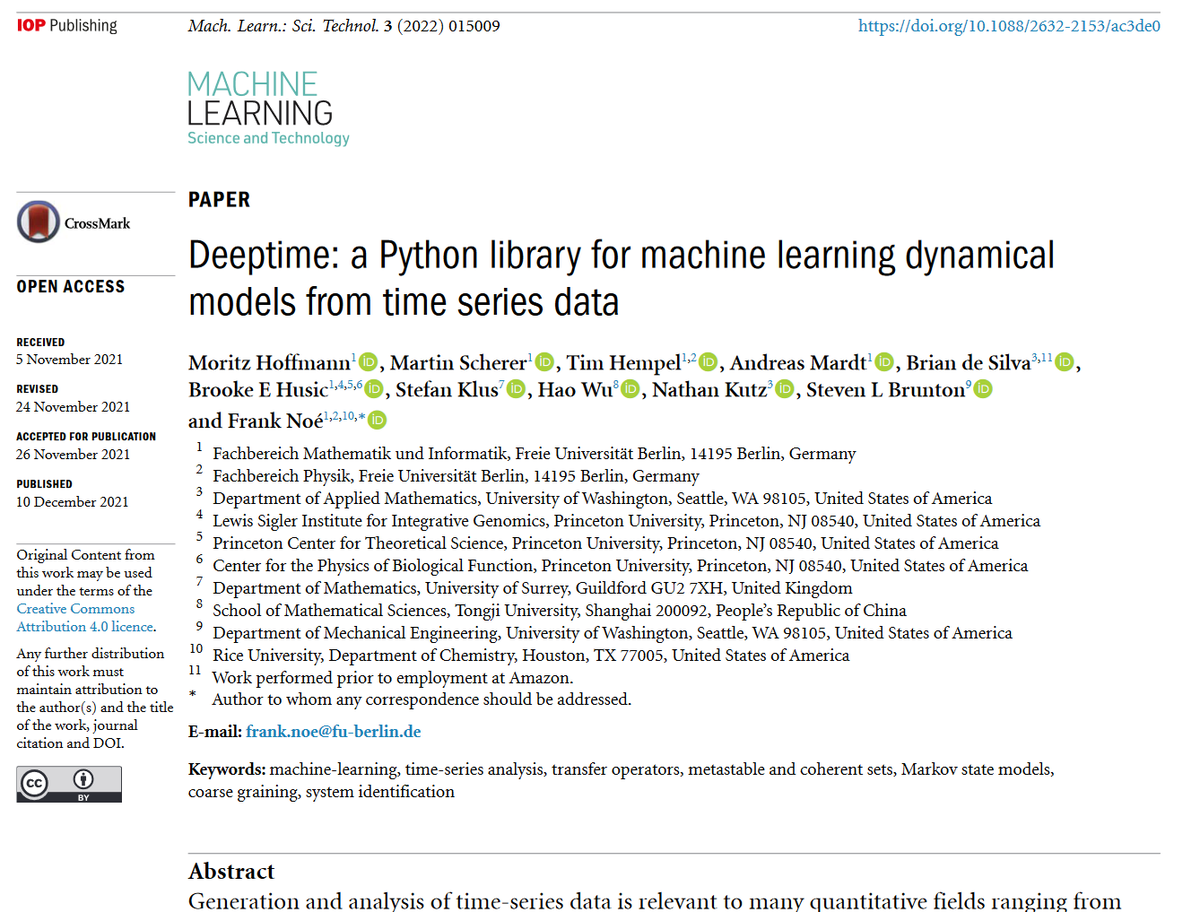


Also Yaoyi Chen welcome to Twitter ;-)



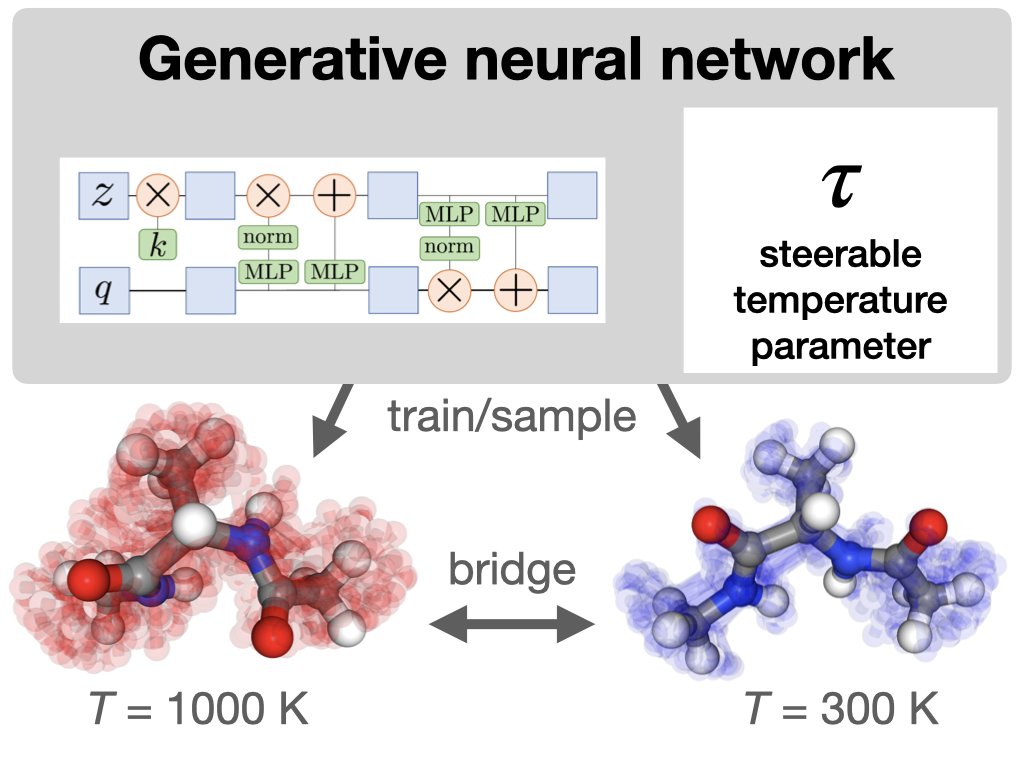
Temperature steerable flows and Boltzmann generators, Manuel Dibak, Leon Klein, Andreas Krämer, and Frank Noé Leon Klein Andreas Krämer Frank Noe Freie Universität #StatisticalPhysics go.aps.org/3MARa93


Check out Michele Invernizzi's new preprint on using #BoltzmannGenerators for #MachineLearning enhanced sampling with replica-exchange molecular dynamics.



With Michele Ceriotti, Gabor Csanyi, and Lixin Sun, we are organizing an awesome conference in Berlin on June 19-23: "Bridging length scales with machine learning: from wavefunctions to thermodynamics". Check it out: sites.google.com/view/cecam-psi…

Another manuscript on the ongoing effort of our labs in the development of accurate coarse-grained models for proteins. With Frank Noe, Yaoyi Chen Andreas Krämer Aleks Durumeric, and Nick Charron. Check it out! arxiv.org/abs/2302.07071


Soon @ ICML Conference ! 🌪️Normalizing flows for rigid bodies and the rotation group SO(3)! 🌪️ We found a way to design smooth normalizing flows for rigid body systems, like ice 👇 Joint work with amazing Michele Invernizzi Pim de Haan and Frank Noe 1/ arxiv.org/abs/2301.11355


Leon Klein introduces flow matching with equivariance. This allows us for the first time to get flows working for iid sampling of molecular structures in Cartesian coordinates in such a way that we can get reasonable Monte Carlo acceptance rates. arxiv.org/abs/2306.15030

🚀 Introducing "Equivariant Flow Matching"! Training equivariant Boltzmann generators has been a challenge, but thanks to a tour de force by Leon Klein, we have an efficient method🌟 On a tight timeframe, Leon executed it with modest supervision and landed it in #NeurIPS2023👏



Hello again 🇺🇸 🏙️ Today was my first day as Machine Learning Researcher at D. E. Shaw Research. I am very excited to be joining an awesome team developing computational tools to transform drug discovery! And it’s unreal that I’ll get to enjoy this view every day now🗽


