PHI-base
@phi_base
This database contains expertly curated molecular and biological information on genes proven to affect the outcome of pathogen-host interactions.
ID: 855384865247821824
http://www.phi-base.org/ 21-04-2017 11:36:58
51 Tweet
88 Takipçi
46 Takip Edilen
















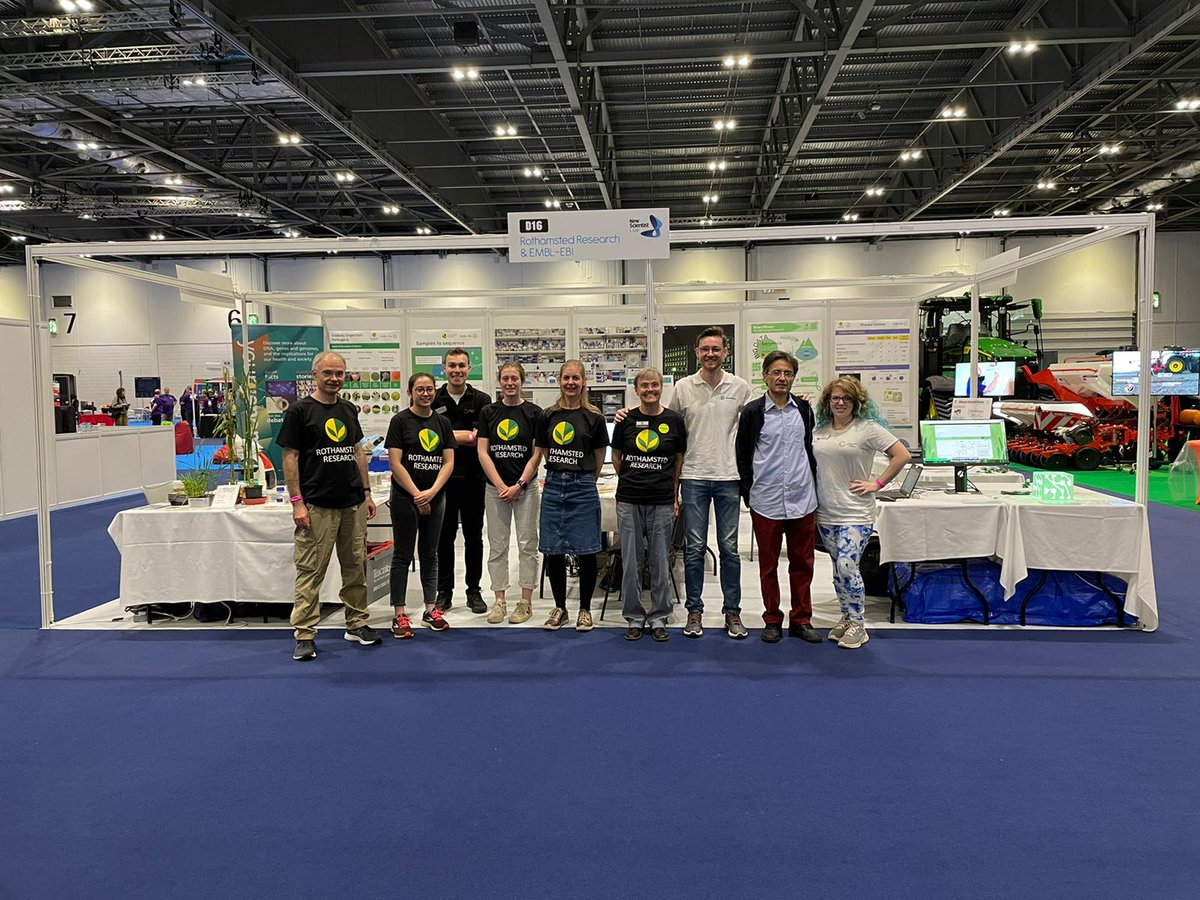
Hello to everyone at New Scientist Live this weekend! Victoria here from Rothamsted Research with a team from KnetMiner EMBL-EBI PHI-base. We’re are at stand D16 talking all about plant pathology and genomes. #OneHealthNSL #planthealth #onehealth #NewScientistLive