
Thomas Plé
@thomas__ple
ID: 1515296582085849088
16-04-2022 11:51:02
16 Tweet
41 Takipçi
40 Takip Edilen

#compchem New preprint: Scalable Hybrid Deep #NeuralNetworks/Polarizable Potentials Biomolecular Simulations including Long-range Effects. Great work by Jaffrelot Inizan Théo Thomas Plé ADJOUA Olivier & collabs: Olexandr Isayev 🇺🇦🇺🇸 Pengyu Ren. EMC2 European Research Council (ERC), #HPC Genci arxiv.org/abs/2207.14276

#compchem New #preprint: Routine Molecular Dynamics Simulations Including Nuclear Quantum Effects: from Force Fields to Machine Learning Potentials. Great work by Thomas Plé. TINKERtools arxiv.org/abs/2212.03137
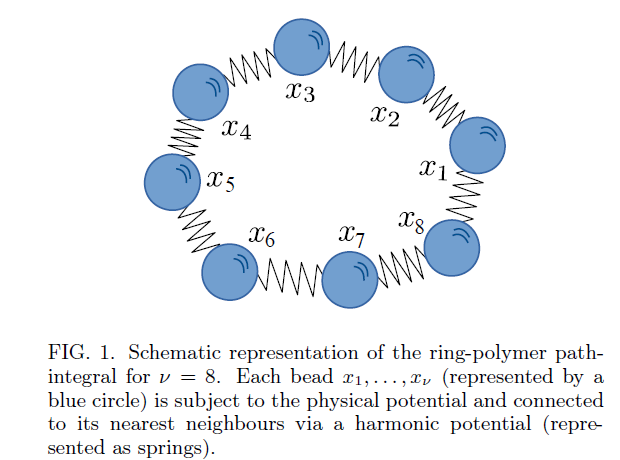

#compchem🚨#preprint🚨:Force-Field-Enhanced Neural Network Interactions: from Local Equivariant Embedding to Atom-in-Molecule properties & long-range effects. Great work by Thomas Plé introducing the hybrid physically-driven #machinelearning FENNIX model arxiv.org/abs/2301.08734
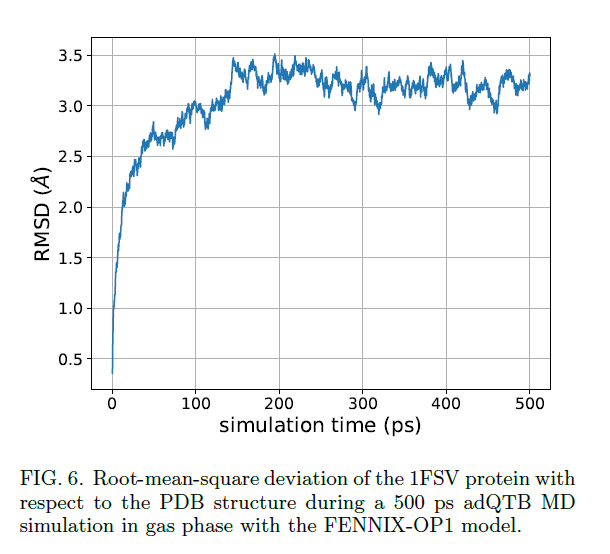

#compchem #MachineLearning #HPC Out Chemical Science. "Scalable Hybrid Deep Neural Networks/Polarizable Potentials Biomolecular Simulations including long-range effects", introducing the Deep-HP #NeuralNetworks multi-#GPU-accelerated platform @Tinkertools doi.org/10.1039/D2SC04…

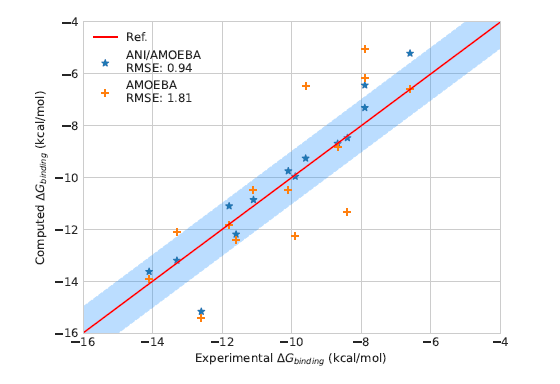
#compchem Preprint:Lambda-ABF: Simplified, Accurate & Cost-effective Alchemical Free Energy Computations. New Colvars #opensource library for NAMD & Tinker-HP. Great efforts: L. Lagardère & L. Maurin & collabs: P. Monmarché & J. Hénin Institut de Biologie Physico-Chimique (IBPC) Paris Laboratoire de Chimie Théorique arxiv.org/abs/2307.08006


🚨Just published in Chemical Science🚨: "Force-Field-Enhanced Neural Network Interactions: from Local Equivariant Embedding to Atom-in-Molecule properties and long-range effects. doi.org/10.1039/D3SC02… Amazing #compchem work by Thomas Plé introducing the hybrid


🥂As the end of year is nearly there, a quick recap on the 2023 the Piquemal Group's papers & preprints in #MachineLearning for #compchem🤖: - Generalized Many-Body Dispersion Correction through Random-phase Approximation for Chemically Accurate Density Functional Theory; J. Phys.
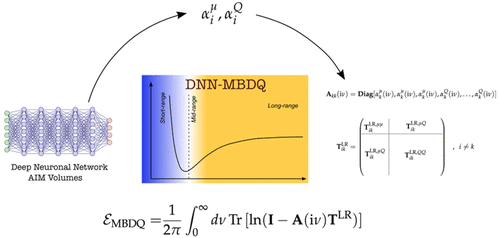



#compchem Just published in JCTC JCIM & JCTC Journals: "Advancing Force Fields Parameterization: A Directed Graph Attention Networks Approach". dx.doi.org/10.1021/acs.jc… In this study, we propose a Graph-Based Force Fields (GB-FFs) model to directly derive parameters for the Generalized
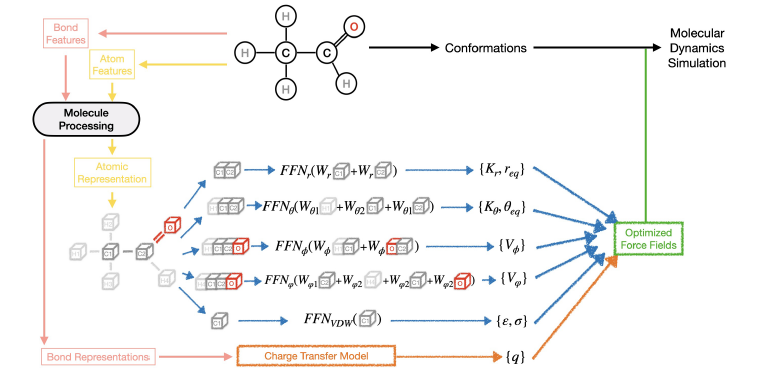

#compchem Happy to see this one out and published The Journal of Chemical Physics as part of the special issue: "Modular and Interoperable Software for Chemical Physics": "FeNNol: an Efficient and Flexible Library for Building Force-field-enhanced Neural Network Potentials. FeNNol is a new


#compchem Just published in Chemical Science: Water-Glycan Interactions Drive the SARS-CoV-2 Spike Dynamics: Insights into Glycan-Gate Control and Camouflage Mechanism. dx.doi.org/10.1039/D4SC04… Towards developing therapeutic strategies against #COVID19, we performed μs-long







#compchem Our paper in J. Phys. Chem. Lett. (The Journal of Physical Chemistry): "Accelerating Molecular Dynamics Simulations with Foundation Neural Network Models using Multiple Time-Step and Distillation" made it to one of the covers! pubs.acs.org/doi/full/10.10…







