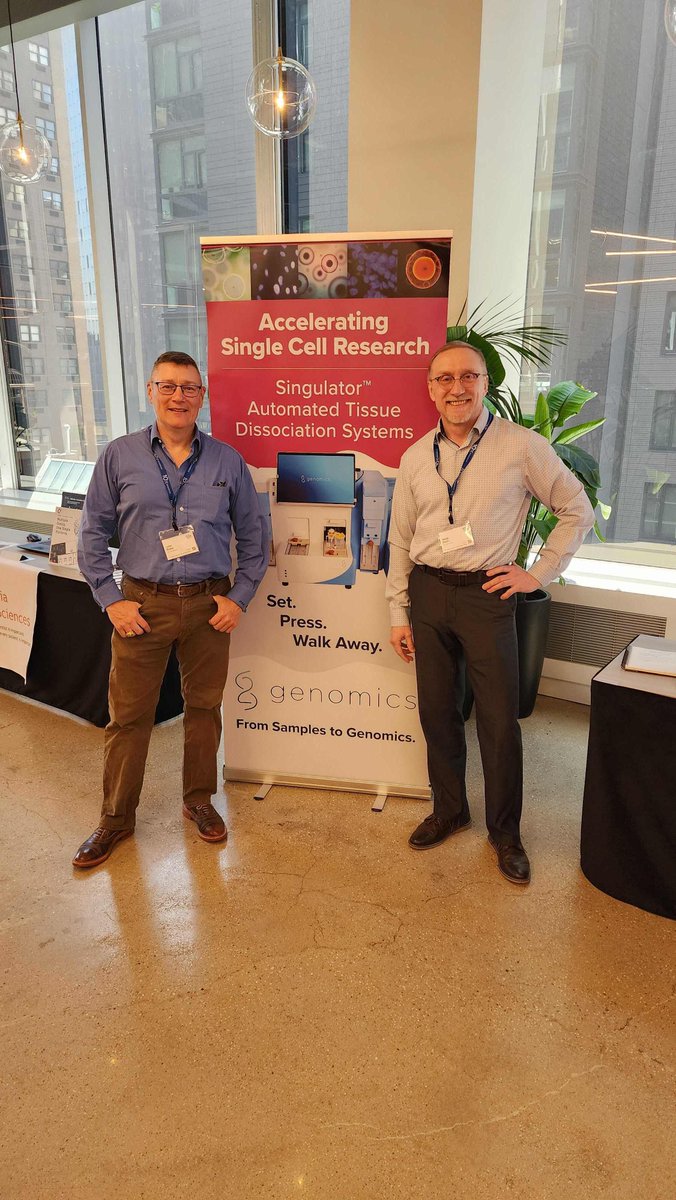
I finally received the FFPE samples from the Ishimoto Lab at Ishimoto Lab at Cancer Institute, JFCR, Tokyo in Japan. It took so long😖
Great thanks to Dr. Nishimura Akiho Nishimura for everything, and I'm also grateful to Dr. Ishimoto for generously providing the precious samples.
#IU #WangLab



I wonder if we will look back and wonder how we did ffpe for so long without normalizing results
#spatialRIN
I get it, researchers work with whatever they get access to. It’s just crazy how little we acknowledge how much sample quality changes results

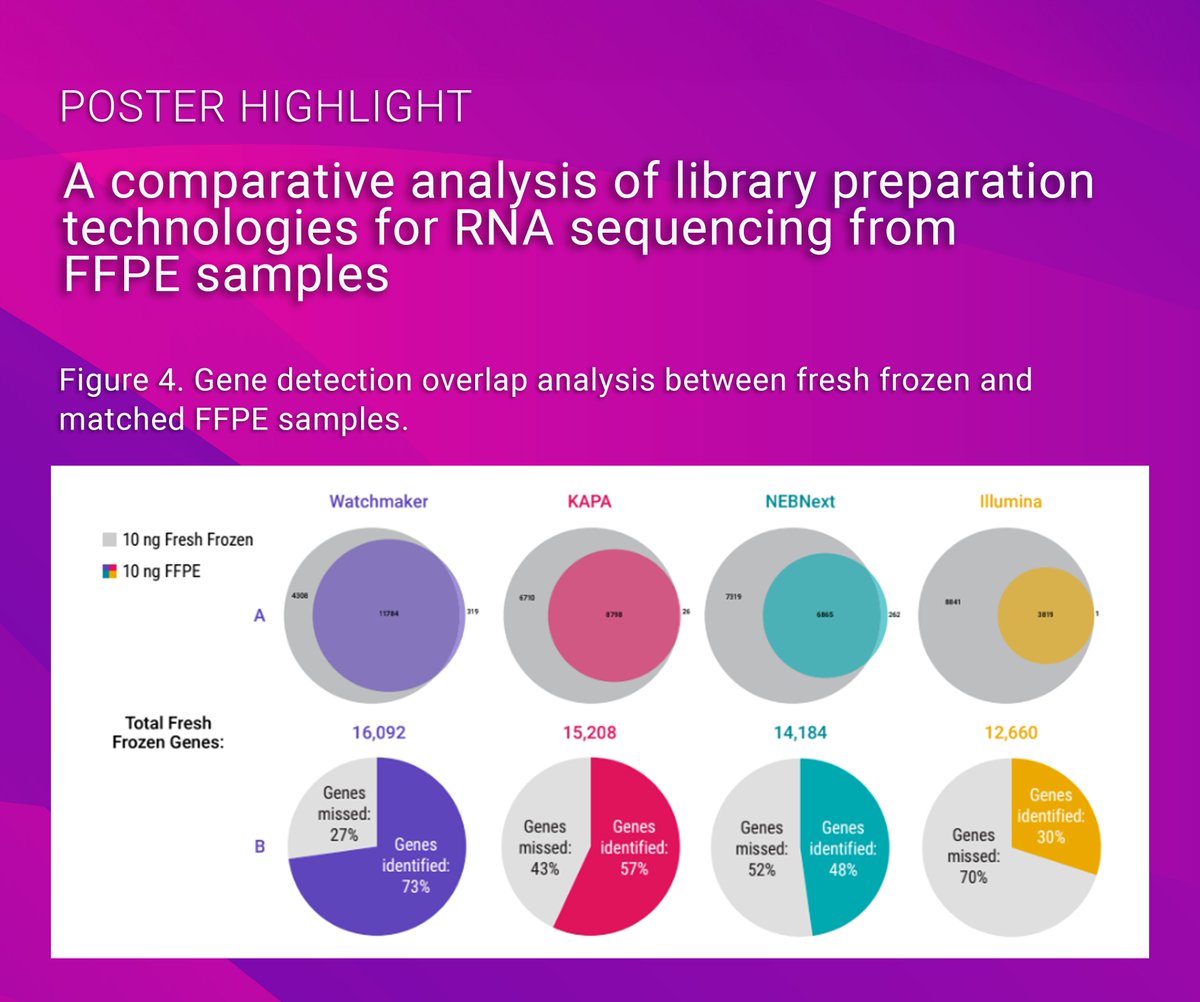
🔍 New scientific poster available! Comparing RNA-seq solutions at AACR 2024 reveals Watchmaker identifies more unique genes & boosts FFPE insights. Maximize data from tough samples! Download here: watchmakergenomics.com/wp-content/upl…
#GetMoreFromFFPE #Genomics #Sequencing



Not everything can be fixed, but your samples certainly can! Did you know that with 10x Genomics you can run #singlecell and #spatialtranscriptomics on fixed samples?
Single Cell #Flex assay allows you to fix and batch samples as you please. It is also compatible with #FFPE …


We couldn't miss #IMSS ! Our resident Yorkshire lad, Russell, will be representing #TeamPreomics in Dublin on May 15, and he's eager to chat to the Irish contingent of #TeamMassSpec about #BeatBox homogenization & making the most of your FFPE tissue samples ow.ly/mqwv50Rut0e


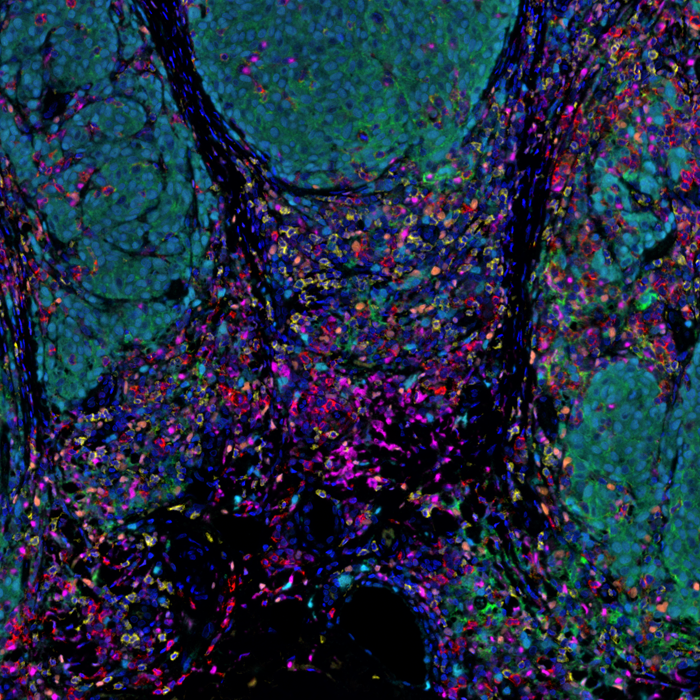
For today's #TissueTuesday , we're sharing a human melanoma FFPE tissue stained w/ a 6-plex MOTiF PD-1/PD-L1 panel auto #melanoma kit & imaged on #PhenoImagerHT .
Learn about our kits crucial for melanoma samples in translational #immunooncology research. bit.ly/3wdqHuc

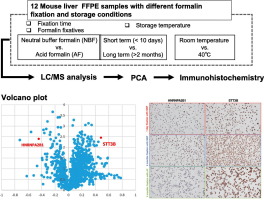
(J Proteom) Proteomic profiling of FFPE specimens: Discovery of HNRNPA2/B1 and STT3B as biomarkers for determining formalin fixation durations dlvr.it/T6bydH (RSS) #MassSpecRSS



Clearing method adapted to FFPE tissues for 3D imaging of nerve fibers, B cells, and tertiary lymphoid structures biorxiv.org/cgi/content/sh… #biorxiv_immuno