Justin Siegel
@siegel_justin
Assoc. Prof. at UC Davis in Chemistry, Biochemistry, and Genomics. Faculty Director of the Innovation Institute for Food and Health. Entrepreneur. Choco Lover
ID: 387666721
http://siegel.ucdavis.edu 09-10-2011 13:19:47
3,3K Tweet
1,1K Takipçi
555 Takip Edilen

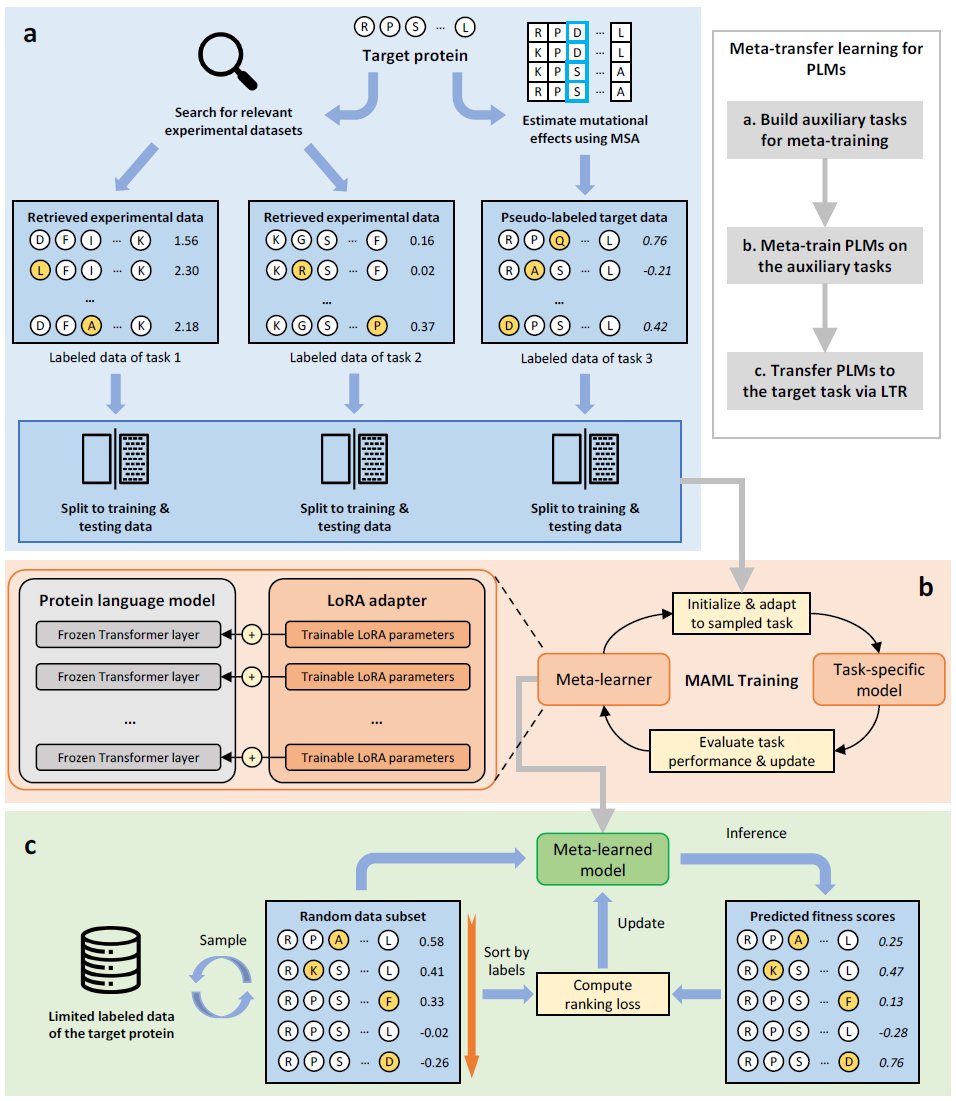
We did 370 experiments to discover that protein language models primarily learn structure and won't scale for protein function prediction. We need new pretraining tasks! Work led by Francesca-Zhoufan Li @ ICLR'25 with Ava Amini Yisong Yue Alex Lu See Alex's thread + the paper for more!



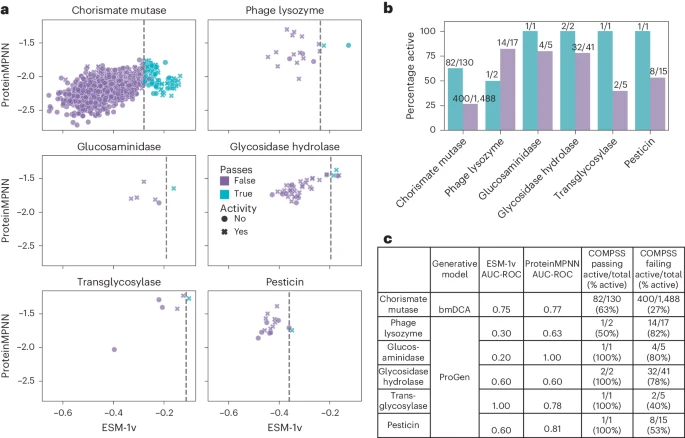
Happy to share the final version of our work describing computational filters for selecting functional ML-generated enzymes in Nature Biotechnology! This was a distributed and remote collab with @TrichomeDoctor Xiaozhi Fu and other members of Aleksej Zelezniak🇺🇦's group. Global Science!



Great work by Kadina Johnston and her team--here is a complete enzyme 4-site dataset for #ML4proteins, folks! #enzyme.

Where our carbon comes from will define our climate future. We developed a synthetic carbon assimilation pathway with new to nature enzymes, enabled by #cellfree synthetic biology. #sustainability Ashty Karim biorxiv.org/content/10.110…

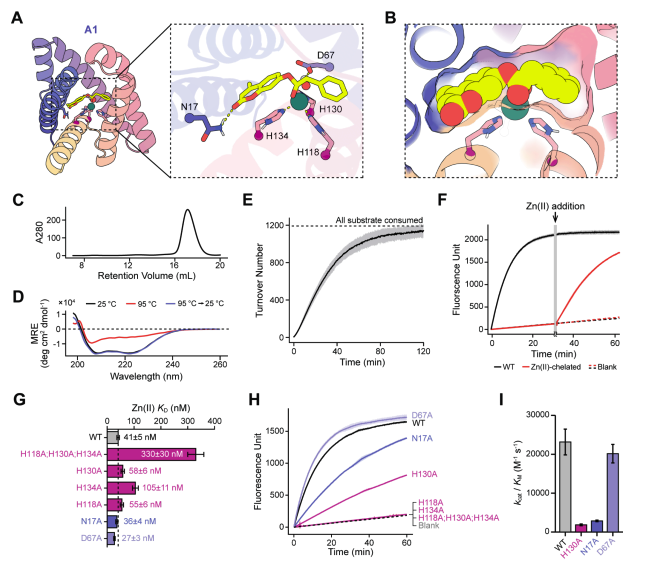
Computational design of serine hydrolases Institute for Protein Design 🚀 New preprints from David Baker! 🚀 1. Groundbreaking study uses advanced computational design to create novel serine hydrolases with unprecedented catalytic efficiencies. These new enzymes could revolutionize


De novo enzyme design!!! | Amazing work from Anna, Sam David Juergens and all the co-authors! Computational design of serine hydrolases biorxiv.org/content/10.110…

Thank you to the Food and Health @ UC Davis for the invitation to share our work, and a big thank you to @joanna_c_chiu Justin Siegel and @AAbrieux! I especially loved the honey tasting, what a treat and great addition to discussions on nutrition and circadian biology!


A couple years ago, Judith Amores convinced me that olfaction was cool, then we convinced Seyone Chithrananda to join us for an internship (he had options!). Hopeful that we can do even cooler things together! Code: github.com/microsoft/olfa… Preprint: biorxiv.org/content/10.110…


We are thrilled to share that Dean H. Rao Unnava has been named Dean of the Year by Poets&Quants, a leading global business school media company. JOHNABYRNE highlights Dean Unnava's innovation and leadership over eight years at #UCDavis: bit.ly/3BteXGl