PathoSense
@pathosense
ID: 1217176080919990274
14-01-2020 20:06:17
19 Tweet
79 Followers
12 Following

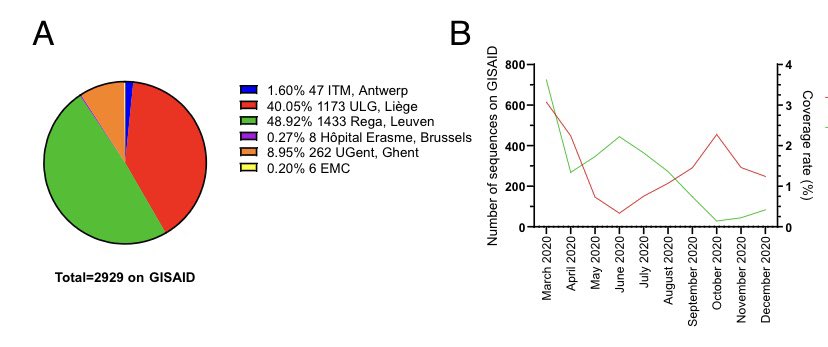
A quick analysis for Belgium to show how we are doing, and what should be improved. This is all based on public data available on GISAID In panel A you see 98% of the genomes were done by either REGA (Piet Maes ), Liège (keith durkin) and Ghent (Sebastiaan Theuns).

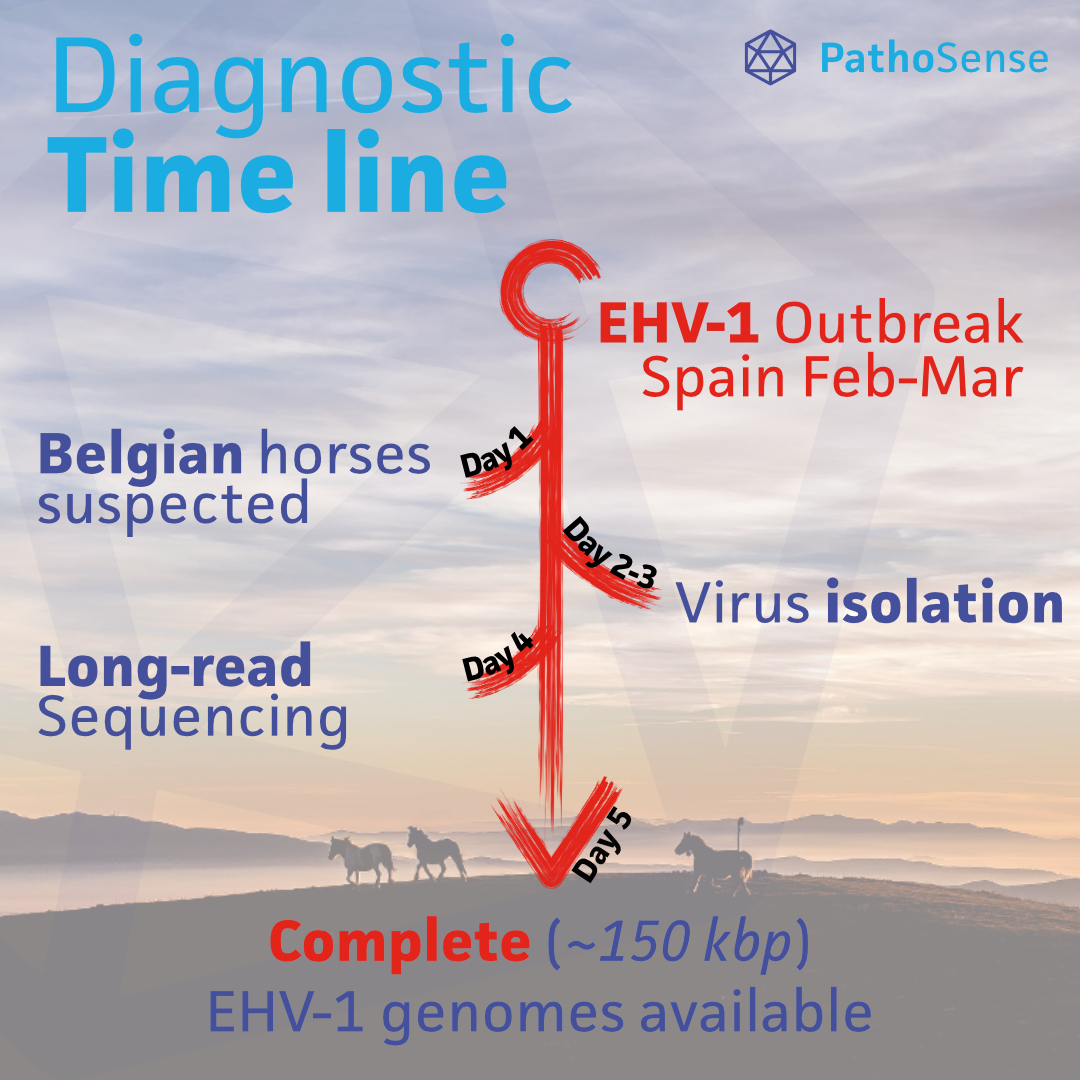
#OutbreakAlert 🐎 February-March: Europe's biggest EHV-1 outbreak was reported in Spain (17 dead & >80 infected 🐎). The #Nauwynck lab isolated EHV-1 from 🇧🇪 suspected 🐎 & by the end of the week PathoSense made complete (150kbp) genomes available with Oxford Nanopore sequencing 1/6


🦠Fresh bacterial release in Microorganisms MDPI Together with Sebastiaan Theuns, PathoSense & UGent VetMed we showed the presence of BSBL-producing Enterobacteriaceae in 💩 of Zoo mammals with Oxford Nanopore sequencing. A potential zoonotic & public health threat? 1/4 bit.ly/3dh8kbD


Delighted to share our #swinedysentery work during the ECCMID meeting, July 2021 🇦🇹. Together with PathoSense & DGZVlaanderen we sequenced 100 Brachyspira hyodysenteriae genomes 🧬 using Oxford Nanopore to unravel #AMR genotypes Vs. phenotypes. #ECCMID2021 Sebastiaan Theuns UGent VetMed


Mycoplasma bovis causes severe respiratory issues & poses a mortality risk to cattle. At #nanoporeconf, Nick Vereecke will demonstrate how nanopore sequencing was used to identify resistance markers & speed up antimicrobial susceptibility analysis. bit.ly/32EQER9


Check out this talk preview from Nick Vereecke. At #nanoporeconf, he'll demonstrate how nanopore sequencing was used to identify resistance markers & speed up antimicrobial susceptibility analysis in veterinary samples 🐄. Register to hear in full: bit.ly/3eHlNc6

Finally out! rdcu.be/cj87M See how we have used long-read metagenomics with Nanopore sequencing to profile a canine fecal microbiome sample and retrieve eight single-contig HQ MAGs 🐕💩🦠🧬 Daniel Pérez @Quinovines Norma Fabregas Olga Francino

Well hello there 💙💚 Let’s finish this Friday with this bacterial 🧬 sequencing run using the Oxford Nanopore Rapid Barcoding Kit! Never thought we would reach these N50 values (> 27kbp) & this caption is only after 30 min sequencing! 🤪🥳 Sebastiaan Theuns #AMR #Diagnostics


Ready for this #nanoporeconf 🧬 today & tomorrow! I am very excited to hear about new Oxford Nanopore stuff that will be released, but also new applications & research from experts in the field! Oh, and don't forget to check my Mini Theatre talk 👇 on Mycoplasma bovis 🐄 PathoSense


New weekend, new #seaofgreen on our MinION. 18h into sequencing and a 10.5 Gbp output so far (still 30h ahead and 83% of Oxford Nanopore pores still sequencing). 👉 RT Bacterial IDx ✅ 🐷 RT Bacterial (Virulence) Typing ✅ ⭕️ RT Bacterial AMR profiling ✅ 🧬 Complete HQC genomes ✅


"Evaluation of Oxford Nanopore sequencing as a diagnostic tool for the rapid identification of Mycoplasma bovis from individual and pooled respiratory tract samples" ⏩ our latest publication in JClinMicro EIC ⏩ A nice collaboration between UGent VetMed & PathoSense 🔽Check the 🧵

A new week, so a new co-first shared publication from the PathoSense lab 🥳 in Microbiology Spectrum "Genome-Wide Association Study Reveals Genetic Markers for Antimicrobial Resistance in Mycoplasma bovis" 🐮 ⏭Read further to learn more in this 🧵 1/6 #sequencing #mycoplasma

📇 The manuscript can be found here: bit.ly/2YC9CcN 🛠 100 Oxford Nanopore High-Quality & Complete bacterial genomes were generated using an optimized M. bovis 🐮 specific Bonito model (Vereecke et al., 2020 BMC Bioinformatics) 📄 2/6 #Bonito Chris Seymour #genomics Kenneth Hoste (mastodon: [email protected])




In conclusion: 🐮 Oxford Nanopore sequencing as rapid new tool to determine acquired AMR & support evaluation of ECOFF values 🐮 current approach allows shortening of the present sample-to-result workflow (IDx & AST) 🐮 future goal = immediate IDx & AST without pre-enrichment 6/6 🧵

Fifty shades of Green 🤢. It seems like Halloween 🎃 👻 is around the corner at the PathoSense lab! Any thoughts which green monster is getting tamed here? 👹👽💀 #labfun #halloween #greenmonster Sebastiaan Theuns #sequencing #phdlife #october #nucleicacids