Geoff Nelson
@geoffmnelson
Oncology bioinformatics @AstraZeneca
ID: 2330663065
06-02-2014 18:09:23
124 Tweet
126 Takipçi
194 Takip Edilen

Check out our new study with @morehouse_ben and the Kranzusch lab looking at how a cognate second messenger induces oligomerization and activation of bacterial STING - fun EM images by Matthew Yip and Alex! nature.com/articles/s4158…

One of the coolest mechanisms we’ve worked on: rdcu.be/cSPbR Matthew Yip and @samanthasedor reveal how the hybrid E2/E3 enzyme UBE2O, which ubiquitinates unassembled ribosomal proteins for degradation, selects its clients.


It’s intuitive that cells will fix aberrant proteins (if possible) before trashing them, but examples remain limited. Our new study by Michael McKenna describes how corrective protein quality control by the P5A-ATPase ATP13A1 prevents wasteful ERAD. authors.elsevier.com/a/1fzWJ_Oylrkq…



Check out the latest paper from the Balasubramanian group with the amazing first author Silvia Galli, where we describe the development of a nanobody to detect and map G-quadruplexes in cells Larry Melidis Siskin @sflynn4 🥳🥳


The latest installment of Luke Coddington studies on how dopamine (DA) shapes learning rdcu.be/c3Ibf This was a tricky one; it combines a few ideas that together argue for a new way to understand DA function and account for data that evaded prior models. 1/n

Brilliant Markus Höpfler presents you the mechanism of autoregulated tubulin mRNA death! The pleasure of having contributed to this story is unmeasurable. Read more about this elegant pathway, and how important it is for a healthy cell division and a healthy human! 👇



So excited to share that my thesis work is out now at Molecular Cell! Karen AdelmanLab and I show that U1 snRNP increases RNA Pol II elongation rate within AT-rich introns, thereby reducing the likelihood of Pol II termination or arrest. Highlights below! authors.elsevier.com/c/1gobY3vVUPNx…

I am excited to share our new preprint! We comprehensively analyze gene expression in male germ cells to reveal transcriptional pausing is required for murine spermatogenesis. Thanks and congrats to my co-authors Dr. Helena Zomer Geoff Nelson AdelmanLab, Irene, Debarun, and Prabu!!

Really excited to share the latest work on #BasalForebrain from our lab: "The behavioral signature of stepwise learning strategy in male rats and its neural correlate in the basal forebrain" buff.ly/44Zmt5j just published in Nature Communications (1/n)


Happy to announce the final version of our paper is now available Nature Communications! nature.com/articles/s4146… Thank you to co-authors Kavya, Geoff Nelson Dr. Helena Zomer, Debarun, Irene, Reza, Huanyu, AdelmanLab and Prabu for a great collaboration of transcription and development expertise!


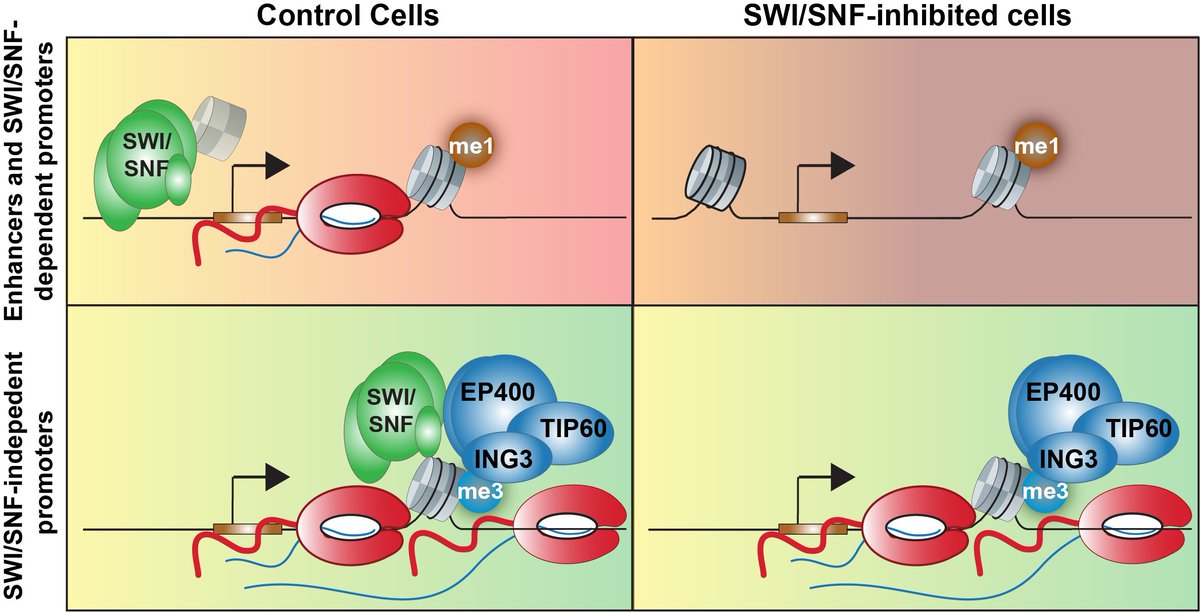
I'm excited to share my manuscript describing our new insights into how Notch signaling induces transcription of target genes. Thanks to BlacklowLab and AdelmanLab and all co-authors! biorxiv.org/content/10.110…