Oxford Nanopore
@nanopore
From portable to population scale, nanopore devices offer direct analysis of DNA/RNA in-real time — towards the analysis of anything by anyone, anywhere.
ID:37732219
http://www.nanoporetech.com 04-05-2009 19:21:00
19,5K Tweets
65,3K Followers
9,9K Following
Follow People

What you're missing matters. Come see for yourself!
At #nanoporeconf , Alexander Wittenberg will discuss constructing high-quality, chromosome scale, fully phased, reference pan-genomes, & breeding new varieties of fungal resistant plants.
Attend online for free: bit.ly/4a4RF5g


We have released some raw Oxford Nanopore data for benchmarking purposes, along with some basecalled outputs.
We use the subset of 20k or 500k for benchmarking tools and methods a LOT!
We thought they would be helpful to others too.
4/5kHz R10.4.1 & RNA004
gentechgp.github.io/gtgseq/


Viruses don’t sleep so why should we? Midnight sequencing protocols 🦠🧬 #H5 #HPAI #Viromics #Alaska #LondonCalling Oxford Nanopore UAA BioSciences ARTIC Network


DNAscent detects BrdU in Oxford Nanopore reads from cultured human cells allowing us to determine DNA replication dynamics on thousands of single sequenced molecules. Fantastic optimisation by Earlham Institute #TransformativeGenomics gave ultra-long reads with N50>120 kb 2/6


Delighted to share our pre-print identifying human DNA replication initiation sites on single molecules by Oxford Nanopore sequencing. Key findings 1/6
biorxiv.org/content/10.110…


Join us at #nanoporeconf Clinical & biopharma day for the 'Rare diseases remagined' panel.
Prof. Joris Vermeesch, Nick Seddon, & Paul Arvidson will unveil next-gen approaches in genomics for early detection & precision therapies. Learn more: bit.ly/3UDPyRd #nanoporeconf


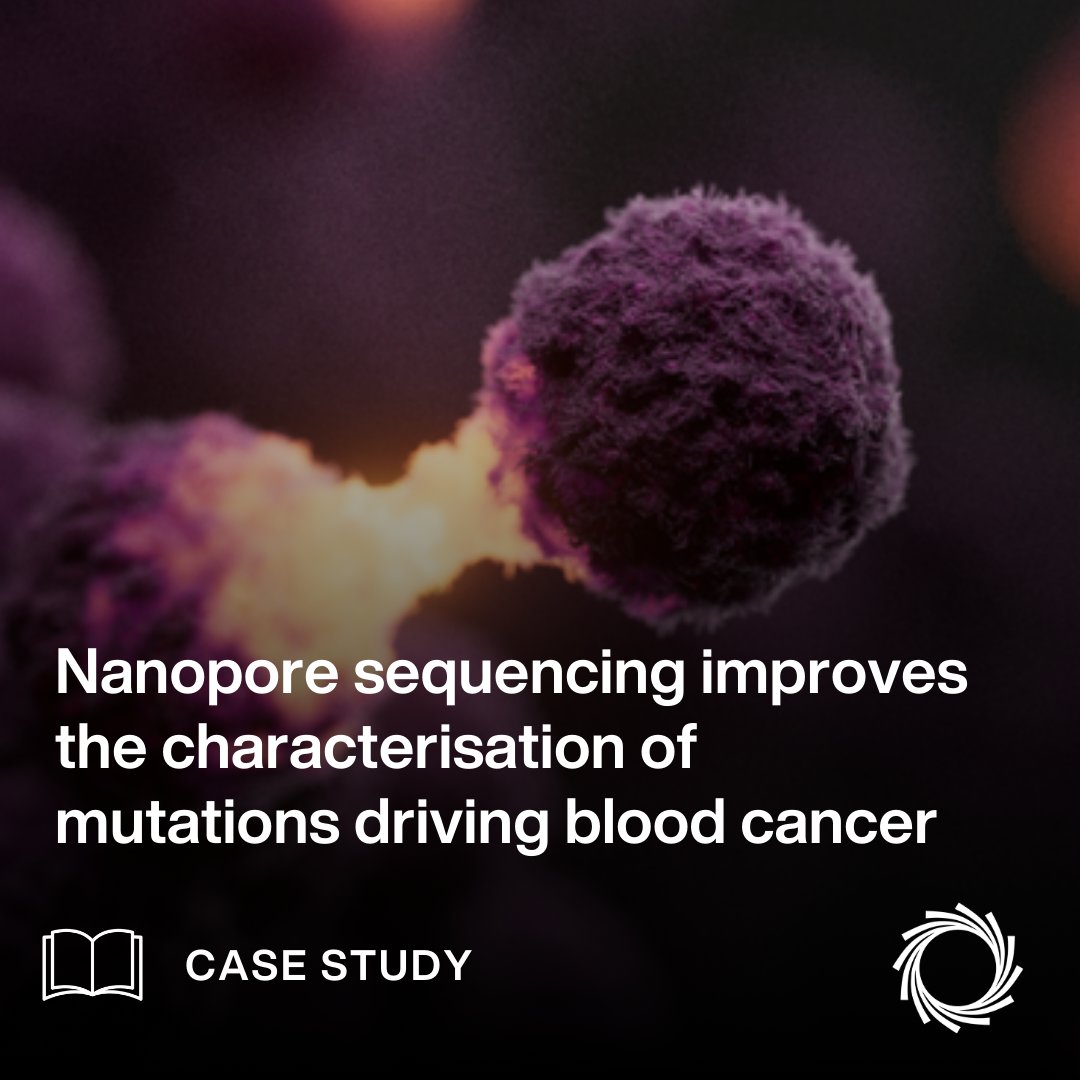
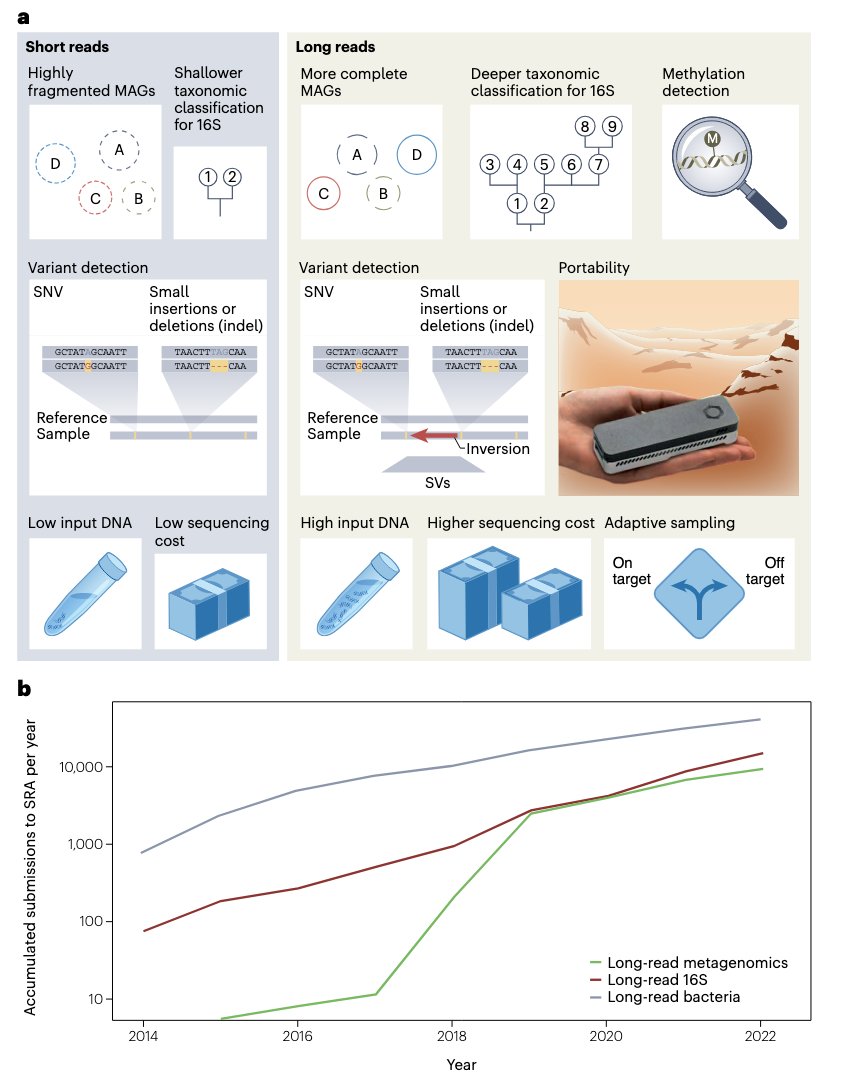
Review paper out today Nature Methods on long-read #sequencing for #metagenomic : rdcu.be/dGeIj Great advancements on utility of Oxford Nanopore PacBio forming novel insights in diversity and resolution. Great work by Daniel Agustinho Yilei Fu Todd J Treangen
BCM HGSC Rice Computer Science

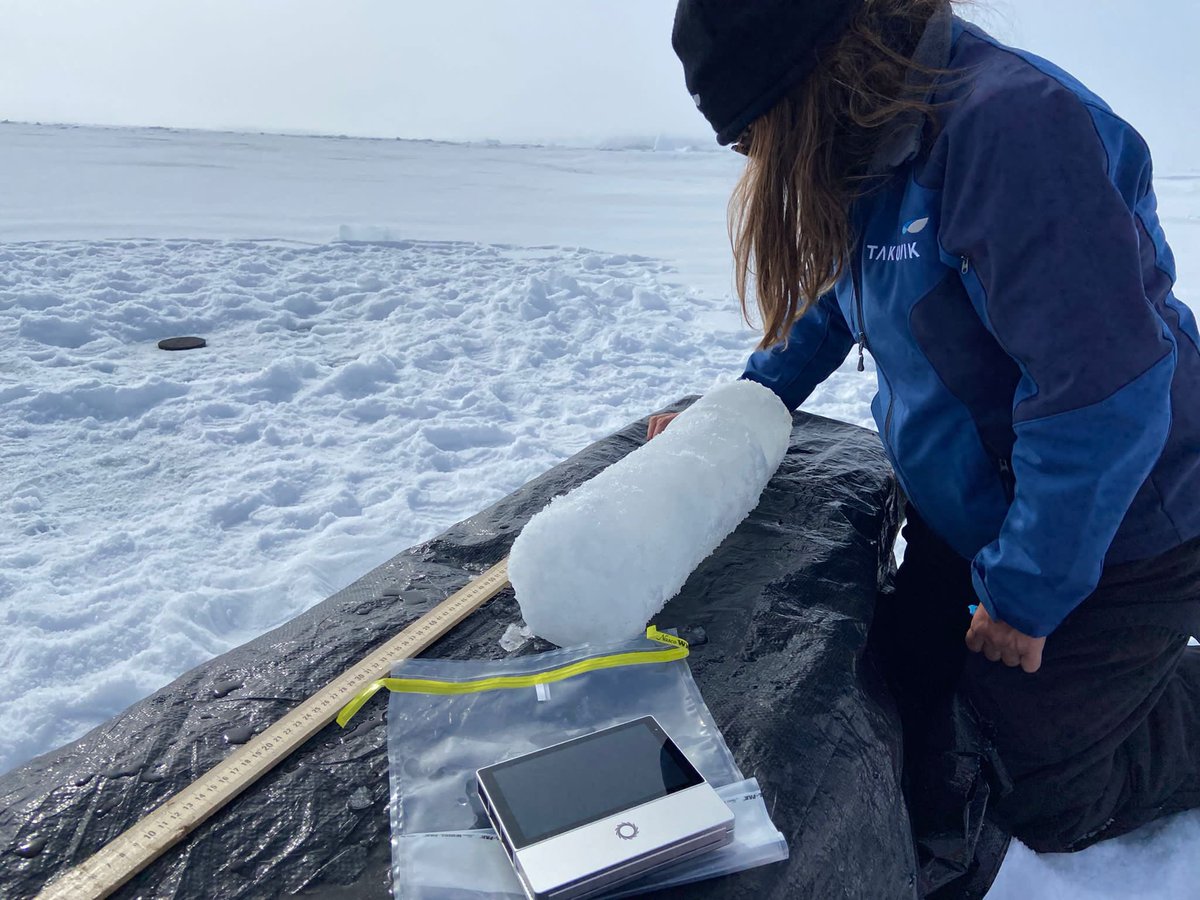
The Acinas Lab's (📷) goal is explore the #microbes that live within the Arctic #Qikiqtarjuaq . They've recovered 26 #metagenomes using the portable MinION device — providing a new insight into this unique microbiome.
Keep up to date here: bit.ly/4aSP672


Direct detection of ribonucleotides embedded in DNA molecules by Oxford Nanopore sequencing, based on ribonucleotides-induced alterations in basecalling features, currents, and dwell times: nature.com/articles/s4200… Genome-Instability&Human-Pathologies Lavinia Grasso

Don't miss this panel on 'Building a rapid, targeted, & responsive infectious disease system' at the #nanoporeconf Clinical & Biopharma Day! Join experts Dr. Inès Hassan, Dr. Chuck Cooper, Dr. Rahul Batra, & Heather Carleton for groundbreaking insights: bit.ly/4aWvUWi


Swapna Uplekar from FIND describing the improvements we’ve made to our Oxford Nanopore AmPORE-TB assay by rapidly implementing World Health Organization (WHO) catalogue version 2.
BDQ sensitivity increased to 69% (WGS 74%)
CLF sensitivity increased to 67% (WGS 78%)
Can’t wait to see the full results 💥

An update from our lab: The first Wales Gene Park double run of sequencing on the PromethION device has started this afternoon. Good to see all that green! Oxford Nanopore Cardiff University #DNASequencing #NGS #longread #10kbprotocol #Promethion

Our latest work on discovering the dark side of CRISPR-Cas9 using our IDMseq Oxford Nanopore sequencing method KAUST Research KAUST Smart-Health Initiative grateful for great collaborators at KAUST Peking University and Altos Labs
link.springer.com/10.1186/s12915…