Hail
@hailgenetics
open-source scalable genetics
ID: 791051969565646848
https://hail.is 25-10-2016 23:01:01
448 Tweet
1,1K Followers
223 Following




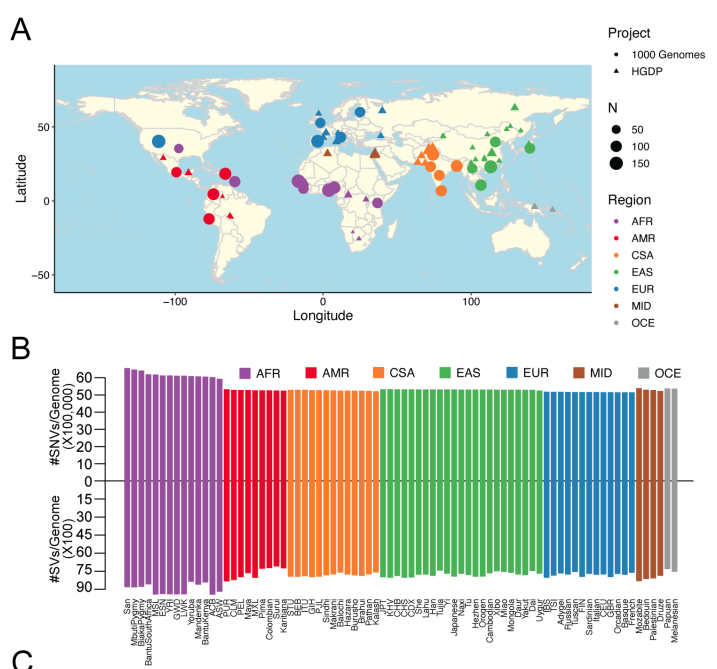
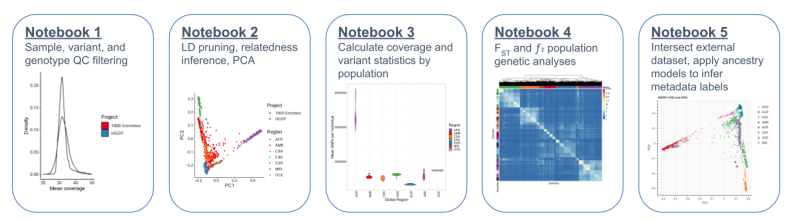
Way to go Zan Koenig and @itsnotmeron for developing scalable cloud-based tutorials for common genomics analyses: github.com/atgu/hgdp_tgp/…. We have also released phased haplotypes for phasing/imputation: gs://gcp-public-data--gnomad/resources/hgdp_1kg/phased_haplotypes

The Broad Institute is accepting applications for its Artist in Residence program! Come hang out with the scientists and make some art! broadinstitute.org/artist-residen…

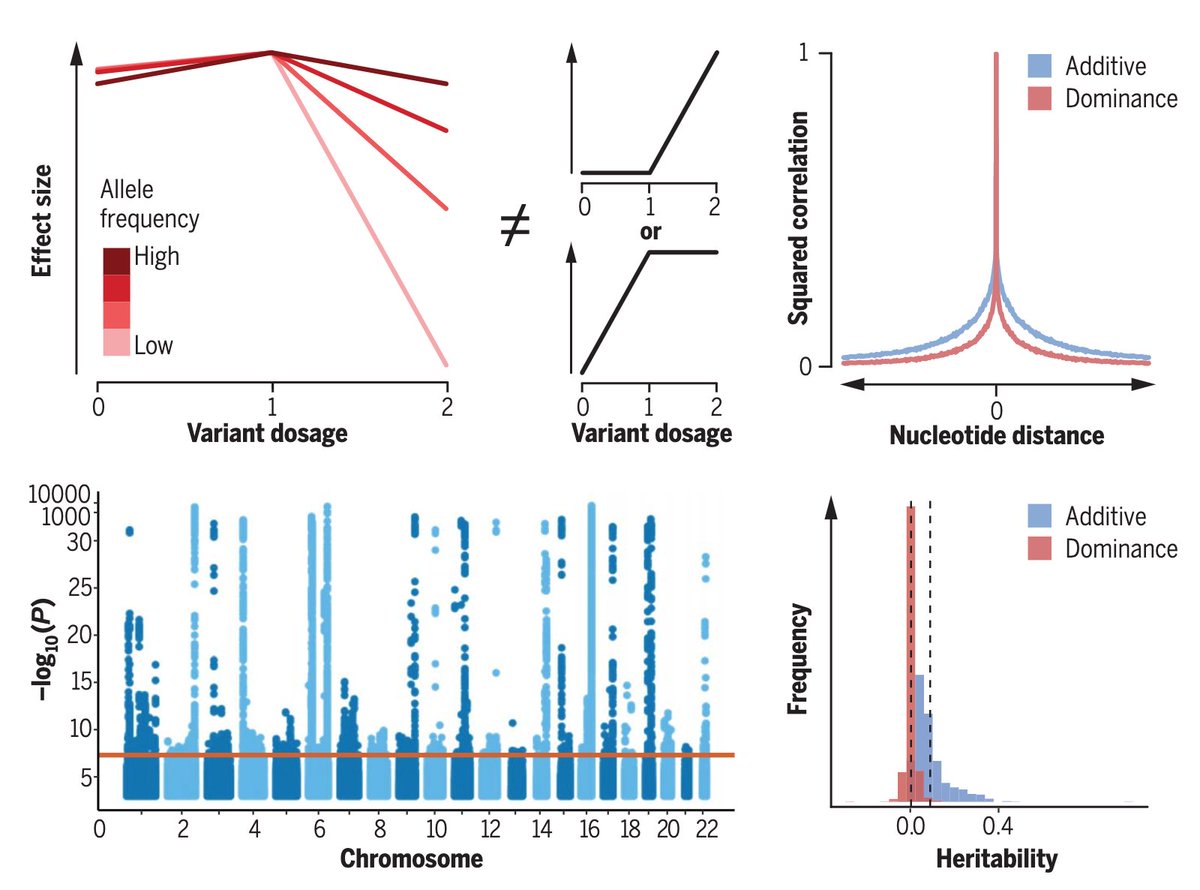
We’re excited to announce that our analysis of genetic dominance in the UK Biobank genotype data is out in Science Magazine (science.org/doi/10.1126/sc…) (1/12)