
ECOGEN Lab
@ecogenlab
Team studying ecological genomics of adaptation in plant communities lead by Fabrice Roux & Fabienne Vailleau @lipme_toulouse @CNRS @INRAE_France @LabEx_TULIP
ID: 1247110650851180544
https://is.gd/xGIB2E 06-04-2020 10:36:32
98 Tweet
319 Followers
54 Following





On recrute un/e ingénieur pour un contrat de 18mois en développement web pour un projet bioinfo #vuejs #genomics Plants Microbes Environment linkedin.com/posts/j%C3%A9r…

I'm excited to announce 3 post doc positions on my European Research Council (ERC) project "Co-evolutionary consequences of biodiversity change" 🌱ecology & evolution of plant-microbe interactions 🌱Soil microbial ecology 🌱Ecological statistics DL 4 December University of Helsinki jobs.helsinki.fi/job/Helsinki-3…




Happy to share this opinion piece on the core microbiota across the green lineage, now open access in: sciencedirect.com/science/articl… Plants Microbes Environment ECOGEN Lab


Mireille Chabaud, from Plants Microbes Environment celebrated her corresponding author manuscript accepted for publication in New Phytol. How #Orobanche cumana penetrates #sunflower root cells to connect to the vascular system. But she will retire April 1st 😭


Thrilled to welcome Ibrahim KEITA as a new PhD student in the team Plants Microbes Environment INRAE Occitanie-Toulouse! Ibrahim will study the regulatory functions of lncRNAs during plant-microbe symbiosis ☘️🧬 a fellowship generously funded by INRAE Santé des Plantes et Environnement and Région Occitanie 🙏


Retour intermédiaire aux agriculteurs du réseau DeepImpact qui nous avaient permis de mener ces 2 campagnes de terrain sur les 200 parcelles en blé et colza


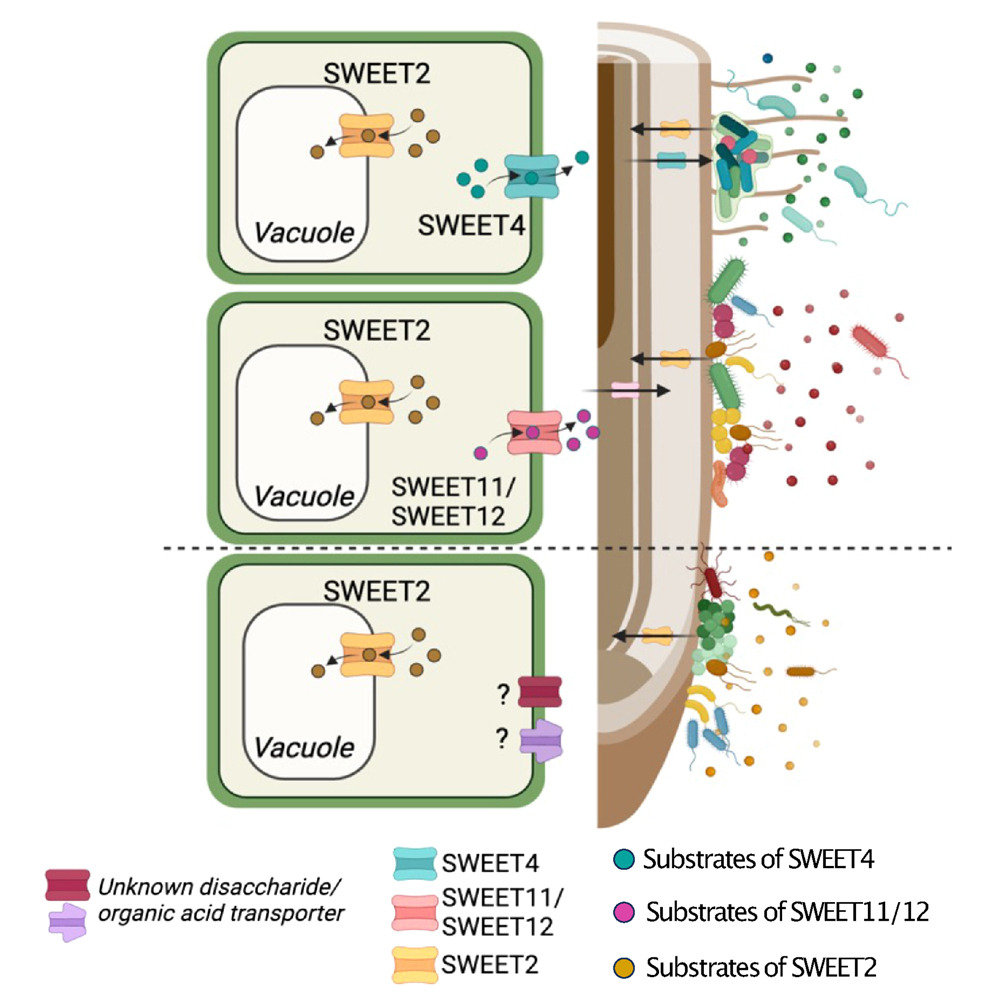
Sugarcoating colonization. Eliza Loo PalomaD &Frommer Heinrich-Heine-Universität Düsseldorf examine organization of #microbiota along longitudinal axis of #Arabidopsis root. SWEET transporters contribute to sugar distribution, which is req'd for spatial colonization of bacteria cell.com/cell-host-micr…

Thrilled to have our collaborative project #SYM2PLANTS selected for funding by INRAE Santé des Plantes et Environnement! With M. Hanemian ECOGEN Lab we will explore how plant-plant interactions impact nitrogen-fixing symbiosis in legumes 🫛🫘☘️ Congratulations to all awardees! spe.intranet.inrae.fr/la-recherche/a…



[#MédiationScientifique] Ravis d'accueillir 70 préparateurs de TP des lycées de l'académie, pour une visite de nos installations #TPMP #transgenèse 🌱 & équipes ECOGEN Lab #lipme_Six 🦠 LIPME_EFIS #lipme_Phyllosym Merci à #FrançoiseJardinaud & Vicedo Céline pour l'organisation
![Plants Microbes Environment (@lipme_toulouse) on Twitter photo [#MédiationScientifique]
Ravis d'accueillir 70 préparateurs de TP des lycées de l'académie, pour une visite de nos installations #TPMP #transgenèse 🌱 & équipes <a href="/EcogenLab/">ECOGEN Lab</a> #lipme_Six 🦠 <a href="/LIPME_EFIS/">LIPME_EFIS</a> #lipme_Phyllosym
Merci à #FrançoiseJardinaud & <a href="/CelineVicedo/">Vicedo Céline</a> pour l'organisation [#MédiationScientifique]
Ravis d'accueillir 70 préparateurs de TP des lycées de l'académie, pour une visite de nos installations #TPMP #transgenèse 🌱 & équipes <a href="/EcogenLab/">ECOGEN Lab</a> #lipme_Six 🦠 <a href="/LIPME_EFIS/">LIPME_EFIS</a> #lipme_Phyllosym
Merci à #FrançoiseJardinaud & <a href="/CelineVicedo/">Vicedo Céline</a> pour l'organisation](https://pbs.twimg.com/media/GRFYDtOWIAAydzz.jpg)


The ECOGEN Lab is searching for 3! PostDocs to join us in the PATHOCOM ERCSyg project in Toulouse, investigating the impact of host and microbial genetics on plant pathogen-pathogen interactions. The links to apply are below. Deadline 29th of Nov!!
![Plants Microbes Environment (@lipme_toulouse) on Twitter photo [#FDS] Le LIPME participe à la Fête de la Science avec 3 ateliers ouverts aux scolaires le 10 Octobre:
☑️Quand plantes🌿et microbes🦠interagissent, c'est sportif !
☑️Athlètes microscopiques
☑️S'adapter,résister,c'est la course pour plantes et microbes
🔗fetedelascience.fr/programme [#FDS] Le LIPME participe à la Fête de la Science avec 3 ateliers ouverts aux scolaires le 10 Octobre:
☑️Quand plantes🌿et microbes🦠interagissent, c'est sportif !
☑️Athlètes microscopiques
☑️S'adapter,résister,c'est la course pour plantes et microbes
🔗fetedelascience.fr/programme](https://pbs.twimg.com/media/F5RbahzXgAADXvD.jpg)

![Plants Microbes Environment (@lipme_toulouse) on Twitter photo [#PhD] 🎓La 3ème doctorante de la semaine nous présente sa thèse sur "Interactions plante-plante: Analyse fonctionnelle d'un gène de réponse à la compétition chez #Arabidopsisthaliana 🌱
Le jury: #ElsaBallini,#PierreAbad, @galaud_jean, #SophieJasinski, #FabriceRoux,<a href="/planticist/">Mathieu H</a> [#PhD] 🎓La 3ème doctorante de la semaine nous présente sa thèse sur "Interactions plante-plante: Analyse fonctionnelle d'un gène de réponse à la compétition chez #Arabidopsisthaliana 🌱
Le jury: #ElsaBallini,#PierreAbad, @galaud_jean, #SophieJasinski, #FabriceRoux,<a href="/planticist/">Mathieu H</a>](https://pbs.twimg.com/media/GO6KBcQWIAEpmWe.jpg)
