Sina Majidian
@sinamajidian
#Bioinformatics | Comparative #genomics | DNA sequencing analysis | PostDoc at @BenLangmead 's Lab @JHUCompSci | Formerly @DessimozLab @UNIL @ISBSIB @WUR
ID: 765158135350890496
https://sinamajidian.github.io 15-08-2016 12:08:10
811 Tweet
454 Followers
785 Following





Looking to analyze allele-specific expression in single cell data? You might want to read this recent review with Guanghao Qi for an overview authors.elsevier.com/c/1jZ~McQbJBL6Q

Want to do a PhD focusing on how complex eukaryotic genomes acquire new genes? Eager to explore how eukaryotic microbes evolve? Come work with us at Laboratory of Microbiology at WUR Wageningen U&R! Apply here: wur.nl/en/vacancy/phd… Share and repost, and contact me if you want to know more!





👀 Extended deadline! Don't forget to submit your abstracts by Sept 6. CSHL Meetings

The Genome Informatics 2024 Meeting will be hosted on the Wellcome Genome Campus Wellcome Connecting Science Learning and Training in Hinxton, UK, November 13-15, 2024. Can Alkan will give an invited talk to feature hardware acceleration of bioinformatics workloads and the BioPIM Project.




📢We are delighted to highlight two news 🥳 1⃣ Our society's members elected ISCB Student Council's next leadership: Pradeep Eranti | @[email protected], Ana Castillo, Wim Cuypers , Sanjana Fatema Chowdhury, Gabriel J. O.-O.. 2⃣ Our current representative to the ISCB News Board of Directors (BoD), Gonzalo Parra, got elected as a




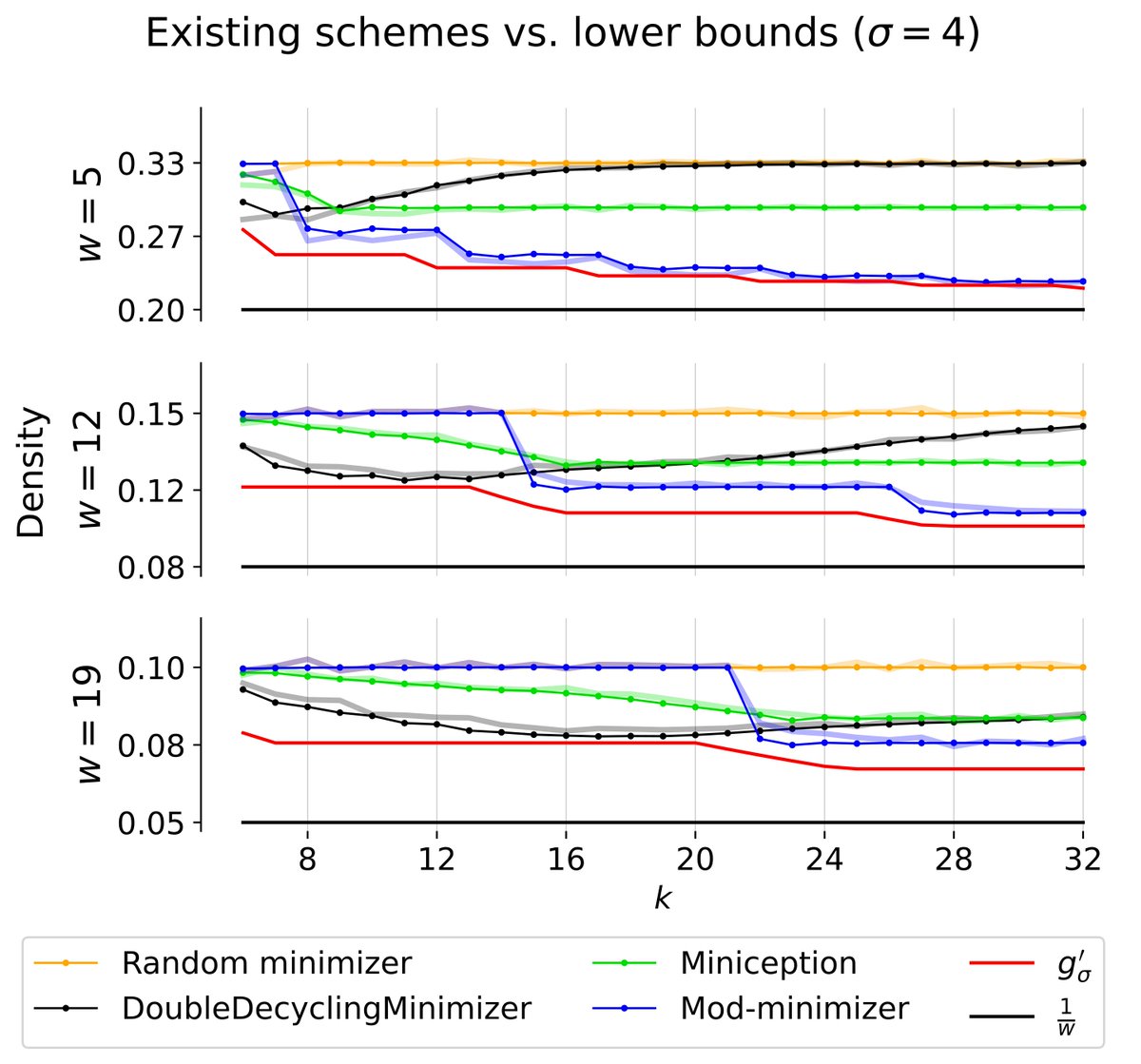
In a new preprint, Ragnar Groot Koerkamp and I led the derivation of a new lower bound, g’ (red), on k-mer sampling scheme density: - our bound appears to be tight for k ≡ 1 (mod w). - existing schemes are nearly optimal! - mod-minimizers are optimal for large sigma for k ≡ 1 (mod w)