Hugging Science
@huggingscience
The @huggingface community effort for science.
ID: 1986824453674979328
07-11-2025 15:54:21
17 Tweet
153 Followers
7 Following


Hugging Face is hosting a “bar crawl for science” at NeurIPS if you’re working on: > autonomous labs > scientific tool use > protein/peptide/etc engineering > materials gen > agents for engineering > fusion and want to meet others, lmk DMs open

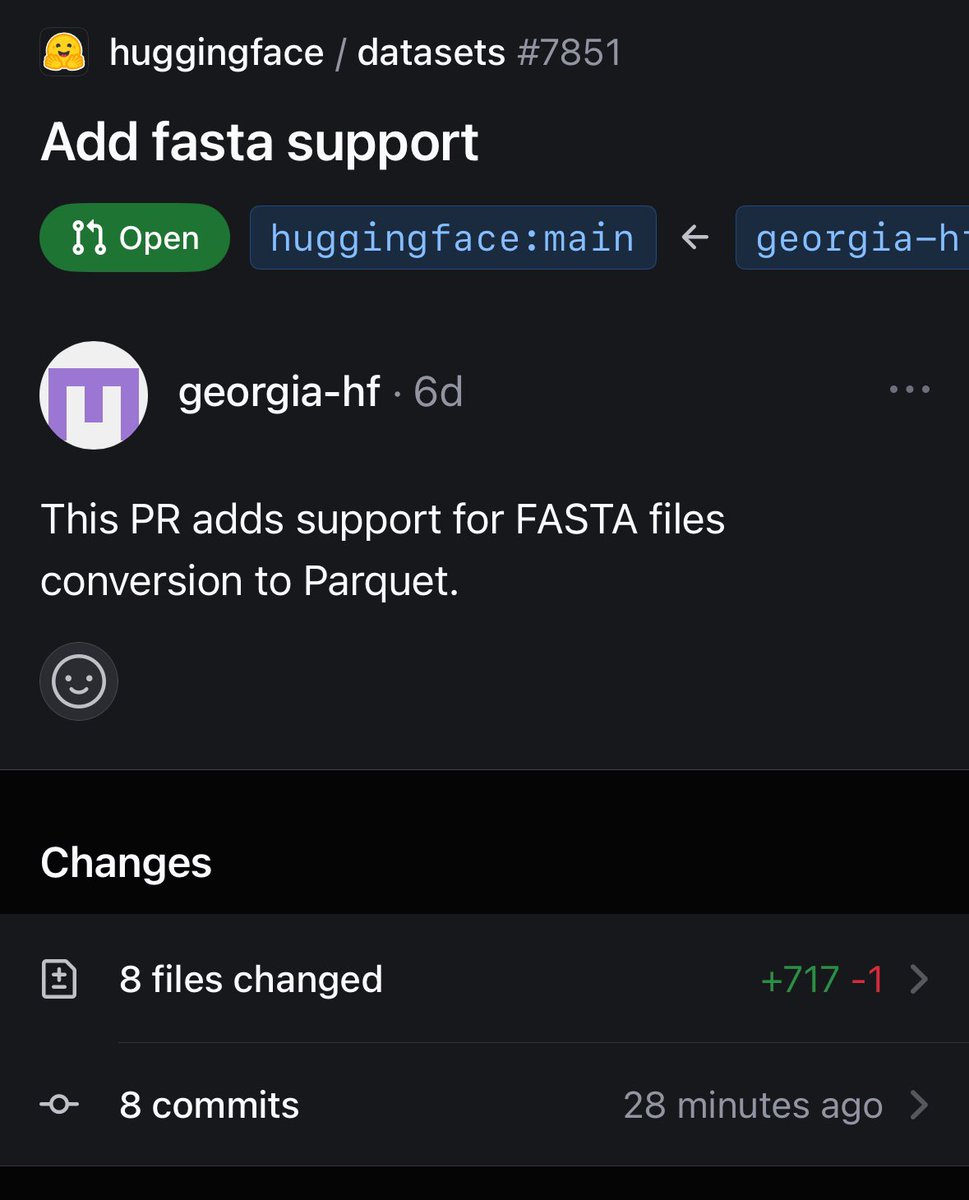
Adding FASTA + FASTQ support for Hugging Face datasets Extending HF to bioinformatics! Need some stuff to test on (especially FASTQ bc PDB has lots of FASTA)—any suggestions? Very happy to test on your data


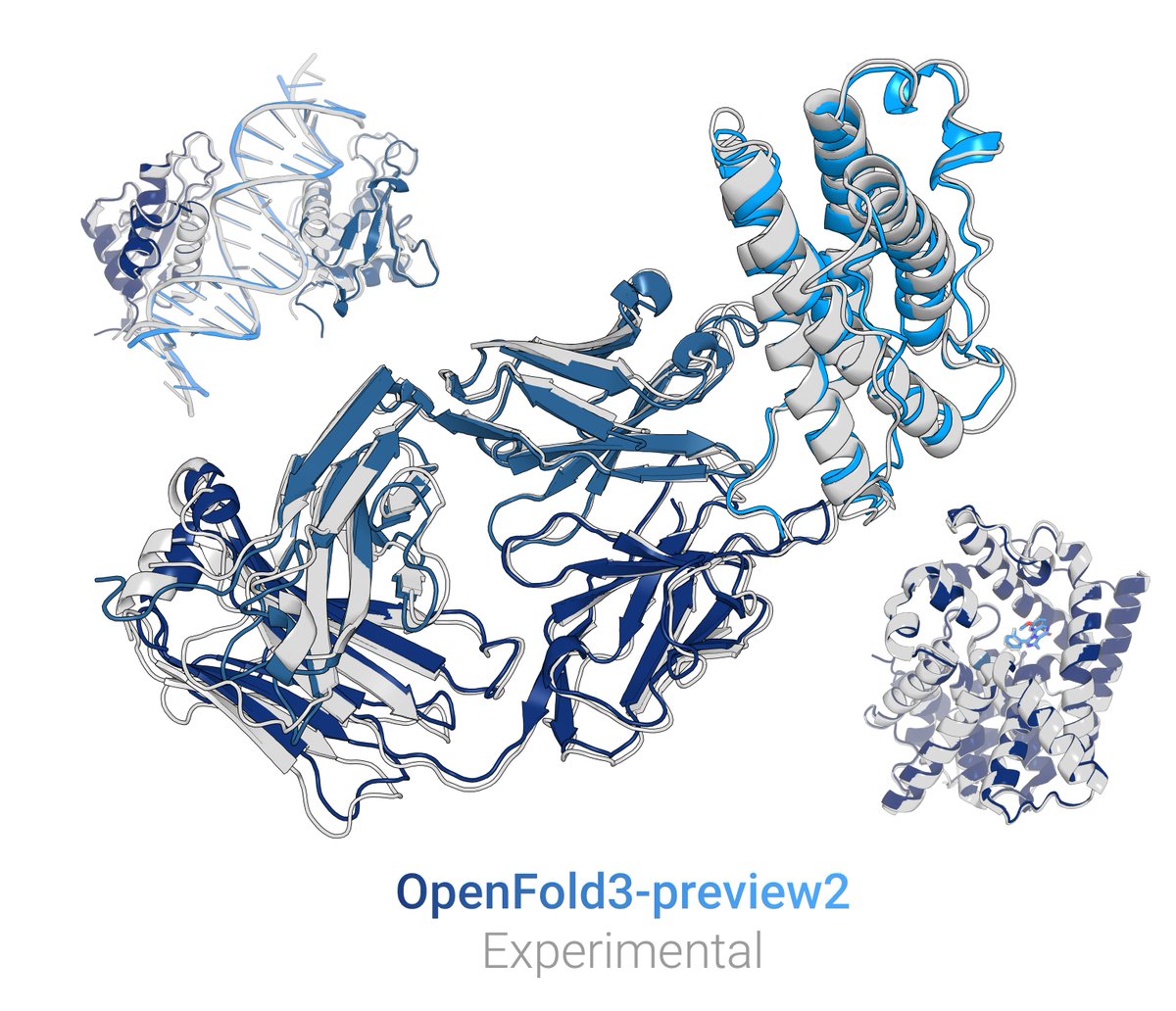
Very cool! The world needs a lot more labs like Gabriel Rocklin 's!


happy to see people enjoying the article, and thanks Georgia Channing for sharing! Go follow Hugging Science!🔬


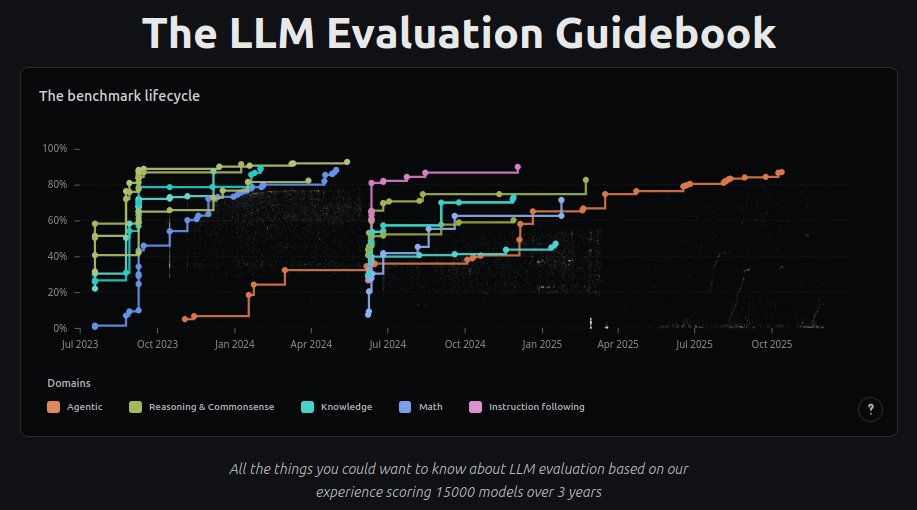
Hey twitter! I'm releasing the LLM Evaluation Guidebook v2! Updated, nicer to read, interactive graphics, etc! huggingface.co/spaces/OpenEva… After this, I'm off: I'm taking a sabbatical to go hike with my dogs :D (back Hugging Face in Dec *2026*) See you all next year!

thanks so much to everyone who came out to the Hugging Science bar takeover last night in San Diego amazing community with epic science coming out of it if you have any photos/videos, plz drop in the thread 🧫🧬🔭🔬💛

Zero code to protein pipeline now on Hugging Science 🤗 As a part of the PDW hackathon, the organizers built inference spaces for: 🧬 RFDiffusion 🧬 RosettaFold3 🧬 BoltzGen2 (+ soon to be MCP servers)

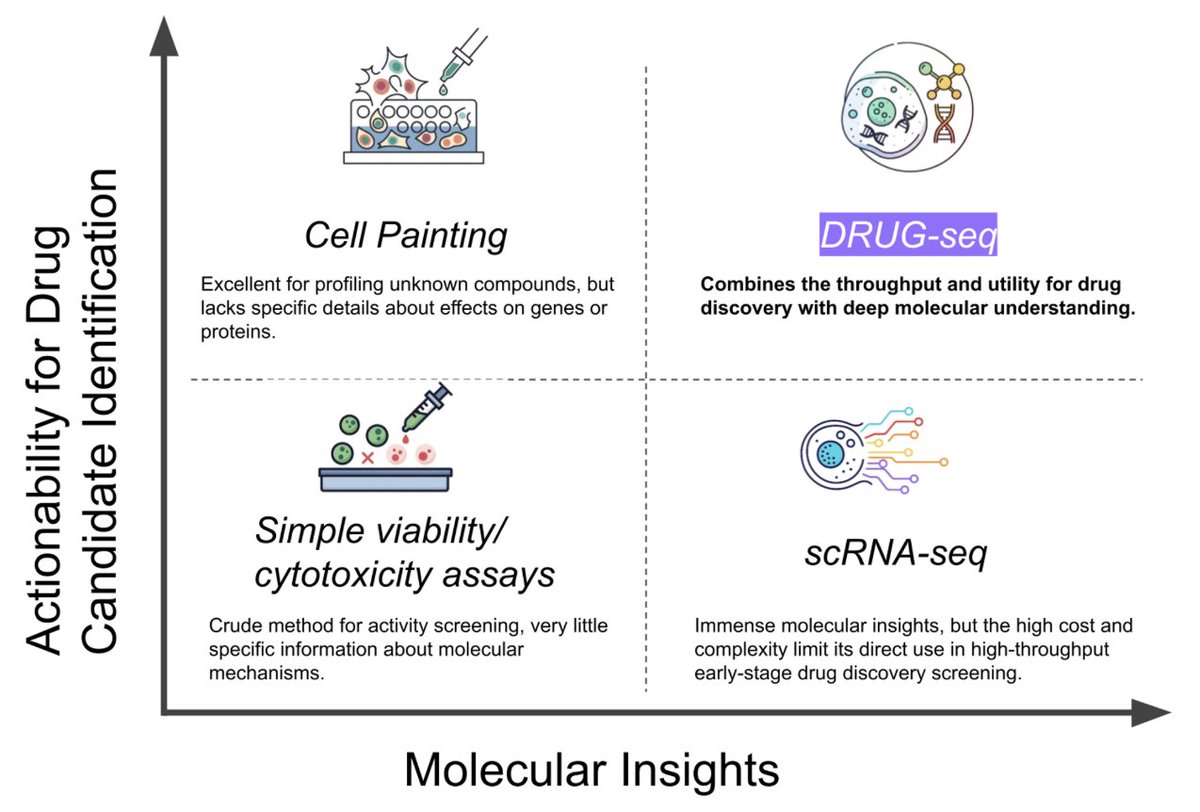
💊💊💊 Ginkgo Bioworks just dropped GDPx4 💊💊💊 29.9 MILLION rows of DRUG-seq data, perfect for benchmarking + large-scale perturbation modeling critical for anyone who can't afford their own high-throughput lab a little more in 🧵

Absolutely killer new blog from one of the OGs in open protein engineering, amelie_schreiber. This is the only piece of writing I've come across that explains to someone in non-specialist language why you need each of these models and what role they all have to play in making sure