Daniel Bryan Goodman
@dbgoodman
Synthetic & comp. biologist @UCSF/@parkerici. Engineering T cells w/ @KoleRoybal & @MarsonLab. DNA synth/multiplex assays/generative models. @geochurch alum.
ID:43357834
http://dbgoodman.com 29-05-2009 16:13:16
2,2K Tweets
1,8K Followers
2,1K Following

How do we interpret molecular mechanisms of protein allostery from deep mut scan experiments? We have built a generalized thermodynamic model to infer and predict how mutations impact inter-and-intradomain allostery. Great collab with Qiang Cui BU
eLife - the journal doi.org/10.7554/eLife.…



Really proud of this one! Used syn bio tools & genome editing to create inducible signaling receptors to remove cytokine dependence for RBC differentiation. Great collaborative work with Porteus Lab Stanford Pediatrics biorxiv.org/content/10.110…

Michael Baym Danielle Beckman Dava Jo Keavney 🏳️⚧️🩻👩🔬 Natalia Quinones-Olvera BioRender Hi Michael - as a BioRender cofounder, wanted to thank you for the feedback and clarify that BioRender does not claim copyright to figures you make, only to the icons/templates made by the BioRender team. We’ll work to fix the wording so it’s much clearer 🙏🏼

Very excited (& nervous) to share what we've been working on the past many years. Only took 3 first authors (Schreuder the experimentalist, Dr Dawn Lin the analyst, and stephen zhang the modeller) & co-senior Weber (computational, modeller, SynBio extraordinaire) biorxiv.org/content/10.110…





Our new study demonstrates extreme ruggedness and high epistasis in the evolutionary landscape of a TF family. Gain/loss function between adjacent nodes, rapid switches in specificity and spontaneous GoF at distinct clades. Colin Jackson Cell Systems
sciencedirect.com/science/articl…


Why can RNA only be linear or circular? It can be branched! We made multi-tailed mRNA, and we had fun! Here is our story led by brilliant graduate student Hongyu Chen: rdcu.be/dB7ck and his cartoon illustration👇MIT Chemistry Broad Institute


LYTACs pioneered by Carolyn Bertozzi opened the door to degradation of cell surface and extracellular proteins through lysosomal targeting. Here, we introduce genetically-encoded LYTACs, or GELYTACs, which are (1) easier to produce by recombinant expression (LYTACs, antibody


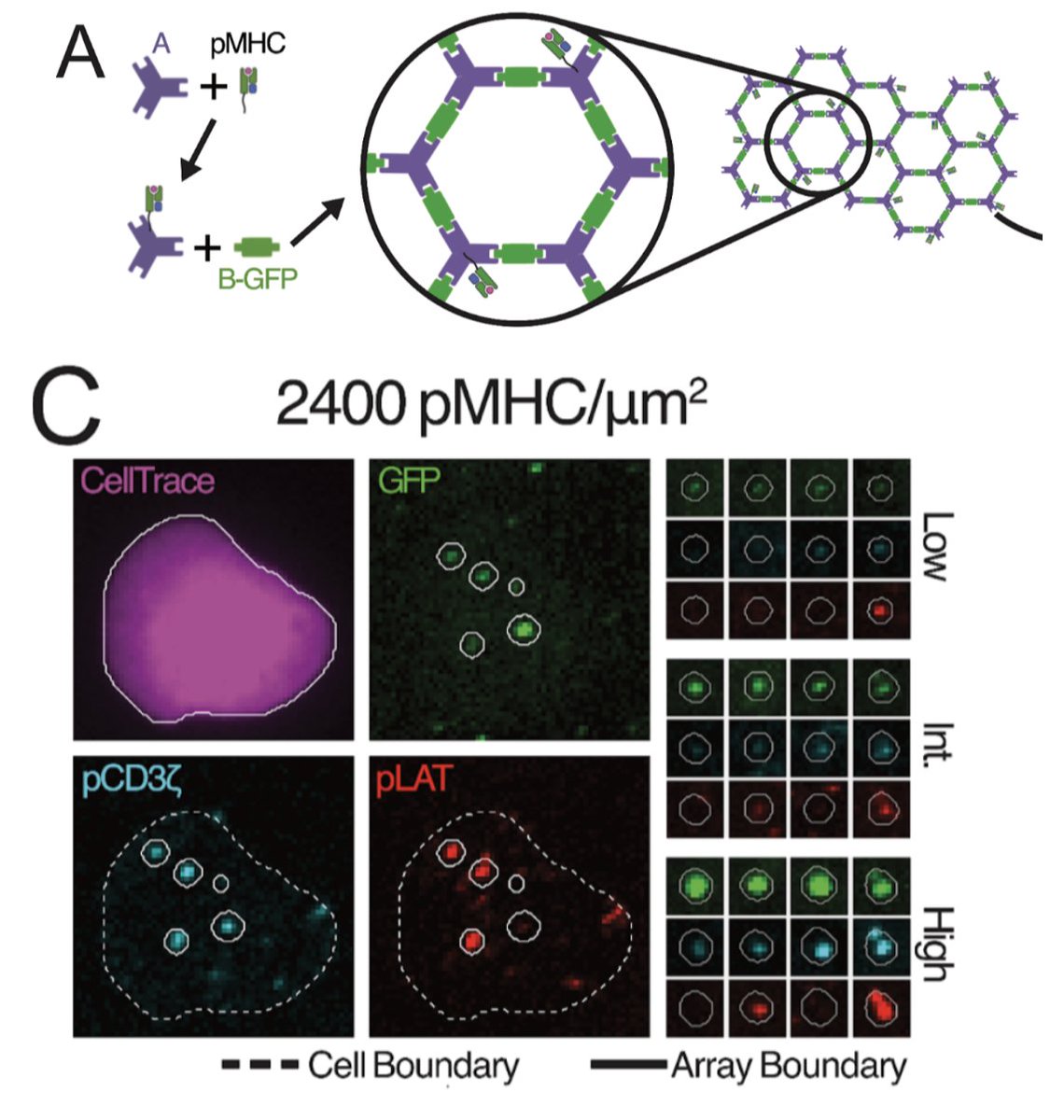
Our latest preprint, on T cell receptor (TCR) signaling by condensate nucleation!
This theory + experiment study was led by the brilliant Will White, with Jay Groves College of Chemistry and David Baker Institute for Protein Design and Ariel Ben-Sasson. (1/n) biorxiv.org/content/10.110…

Ever wonder how microbes interact with the host at a molecular level? Check out our latest paper in @nature with Noah Palm and Curtis Huttenhower where we developed a host:microbe profiling technology, BASEHIT, to atlas interactions between commensal microbes and the human exoproteome.