Fool
@methylrot
Code 🐒 Lab 🐁
ID: 979834277780185088
30-03-2018 21:34:31
115 Tweet
10 Takipçi
221 Takip Edilen

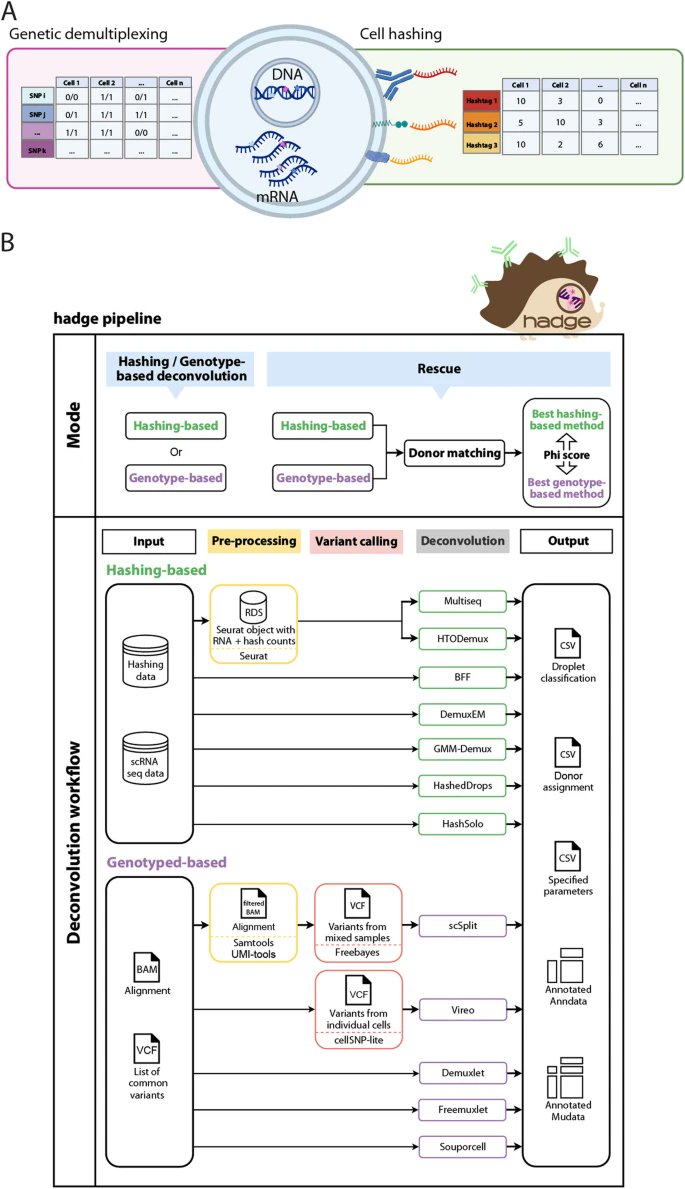
Excited to share with the single-cell community the most recent work of Tomàs Montserrat Ayuso and AnnaEsteveCodina from CNAG, something we think will be a game-changer in #scRNAseq quality control: biorxiv.org/content/10.110… bioRxiv Get ready for the tweetorial! 1/10





ICYMI SciPyConf... quak 🦆 is a scalable, interactive data table built with #anywidget - 🖱️ crossfilter & sort millions of rows in real time - 🔄 view any ApacheArrow __dataframe__ - ⚡ powered by Interactive Data Lab mosaic & DuckDB - 📓 materialize sub-views back in Project Jupyter




Earlier this week we carried out a preliminary analysis of our first 10x Genomics Xenium prime (5,000 gene) run. We had designed this experiment to get a sense of how the prime chemistry performed compared to the Xenium V1 chemistry.