Alessandro Palma
@ale__palmaa
PhD student at @HelmholtzMunich and @TU_Muenchen | ML and computational biology
ID: 1538212324703719426
18-06-2022 17:29:43
27 Tweet
198 Followers
365 Following


Mixed Models with Multiple Instance Learning (MixMIL) received an Oral & Outstanding Student Paper award at AISTATS Conference last week! 🏆 MixMIL enables accurate & interpretable patient label prediction from single-cell data by adding attention to GLMMs.#singlecell #MachineLearning


We introduce Spatio-Spectral GNNs (S²GNNs) – an effective modeling paradigm via the synergy of spatially and spectrally parametrized graph conv. S²GNNs generalize the spatial + FFT conv. of State Space Models like H3/Hyena. Joint work w/ Arthur Kosmala Daniel Herbst Stephan Günnemann


Deep learning with differential privacy can protect sensitive information of individuals. But what about groups of multiple users? We answer this question in our #NeurIPS2024 paper arxiv.org/abs/2403.04867 Joint work w/ Mihail Stoian Arthur Kosmala Stephan Günnemann. #Neurips (1/7)



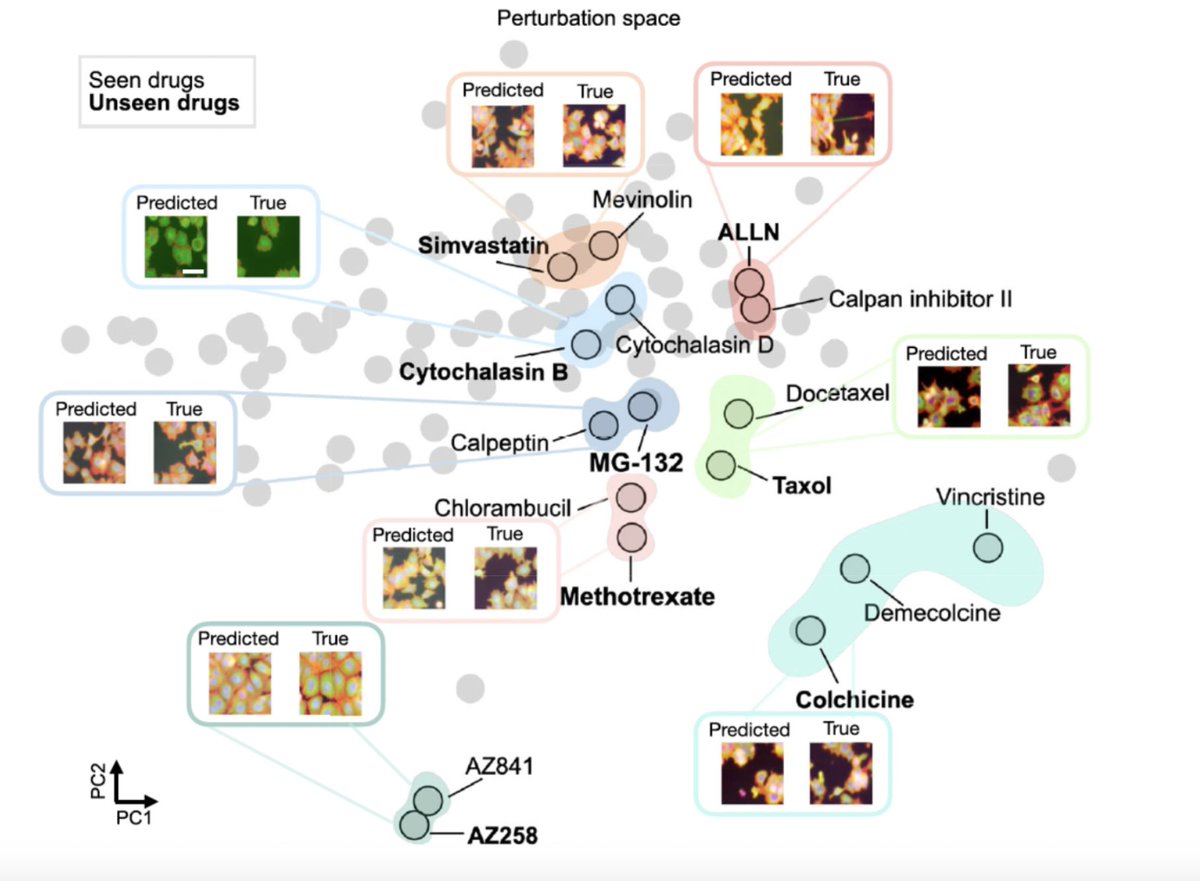
(1/8) IMPA is published now in Nature Communications. (a) It can generate phenotypic cell painting/microscopy data images under unseen drug and genetic perturbations. (b) It learns a perturbation map of treatment similarities (small molecules together with genetic) and (c) it corrects

To figure out what it takes to make flow matching models solve single-cell biology questions, I discussed with Alessandro Palma, Doron Haviv, and Lazar Atanackovic. The insights they shared are presented here: tinyurl.com/278s5m58 Alessandro Palma Doron Haviv Lazar Atanackovic

Thrilled to announce that we just presented „MAGNet: Motif-Agnostic Generation of Molecules from Scaffolds“ at #ICLR2025 🧲 Johanna Sommer Bastian Grossenbacher-Rieck Fabian Theis Stephan Günnemann For those who couldn’t make it to our spotlight: openreview.net/forum?id=5FXKg…



Happy to be in Boston to present nf-core/tumourevo pipeline, a joint effort from all the Cancer Data Science Lab at Università di Trieste! Thank to our PI Giulio Caravagna and Nicola Calonaci for the supervision and also to my colleagues for the effort! Check out at nf-co.re/tumourevo/dev/

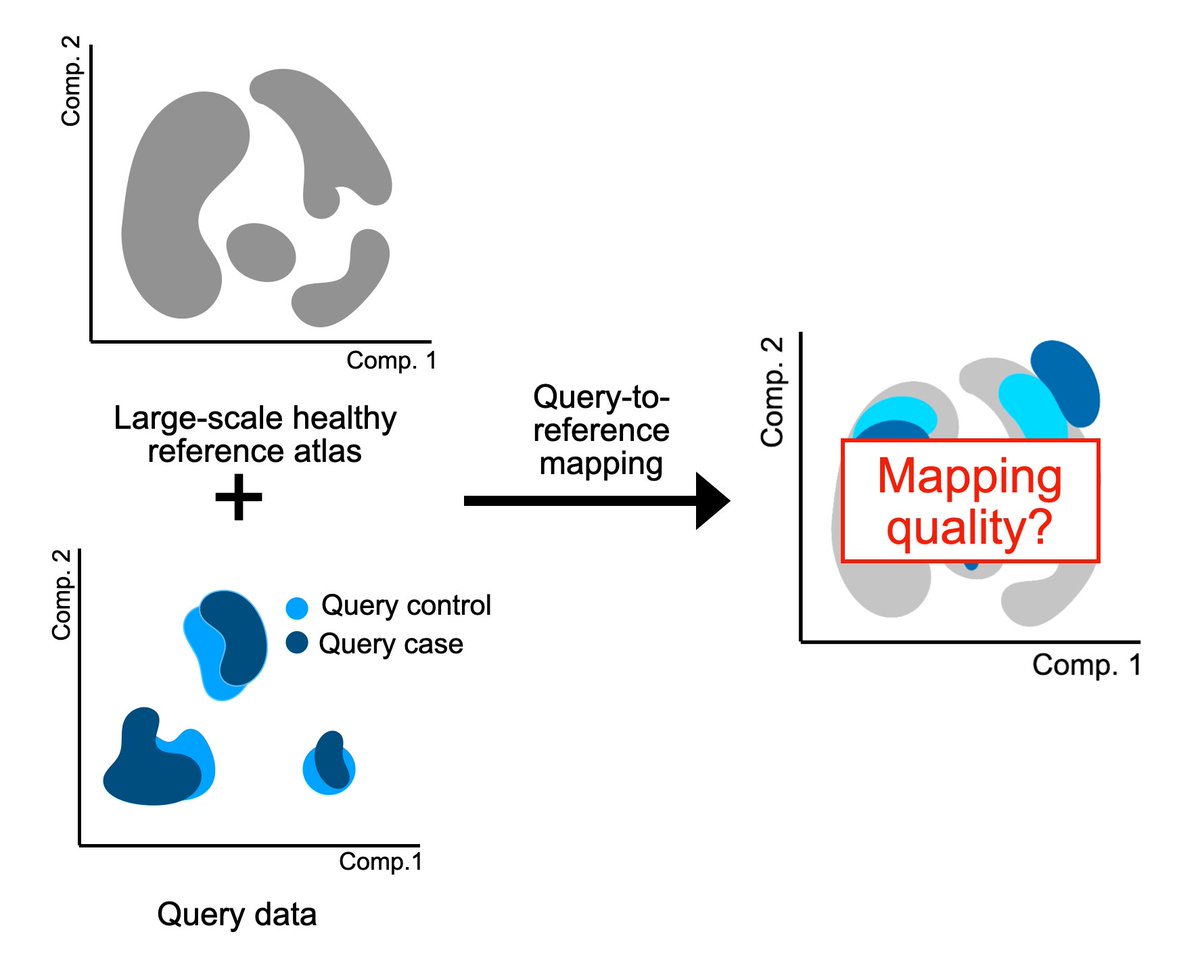
Analyzing your single-cell data by mapping to a reference atlas? Then how do you know the mapping actually worked, and you’re not analyzing mapping-induced artifacts? We developed mapQC, a mapping evaluation tool biorxiv.org/content/10.110… from the Fabian Theis lab. Let’s dive in🧵

How private is DP-SGD for self-supervised training on sequences? Our #ICML2025 spotlight shows that it can be very private—if you parameterize it right! 📜arxiv.org/abs/2502.02410 #icml Joint work w/ M. Dalirrooyfard, J. Guzelkabaagac, A. Schneider, Y. Nevmyvaka, Stephan Günnemann 1/6

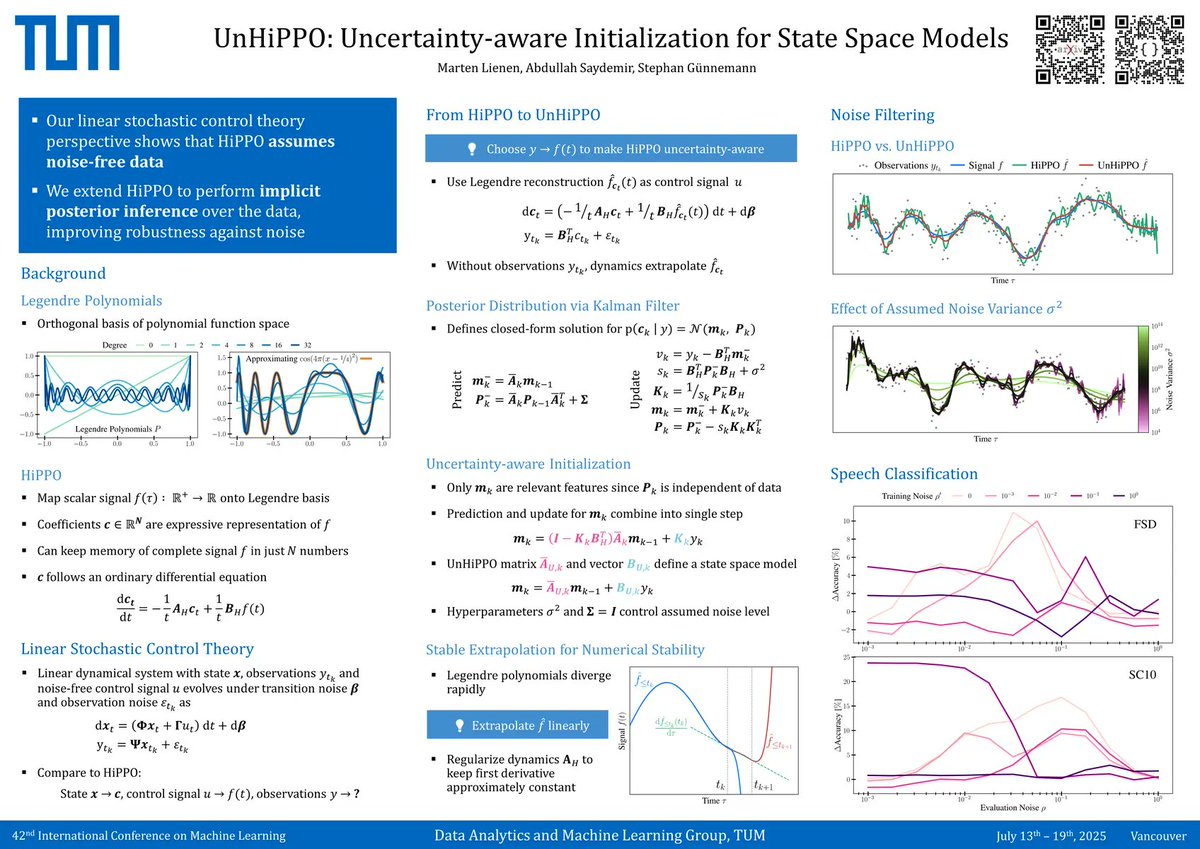
Real data is noisy but HiPPO assumes it's clean. Our UnHiPPO initialization resists noise with implicit Kalman filtering and makes SSMs robust without architecture changes. #ICML poster: Thu 11am E-2409 Paper: openreview.net/forum?id=U8GUm… Code: github.com/martenlienen/u… w/ Stephan Günnemann