
Jan Leipert
@pingvinel
PhD student @kieluni, MS-based proteomics and microfluidics for sample preparation
ID: 912010934658494466
https://www.iem.uni-kiel.de/de/systematische-proteomics 24-09-2017 17:48:45
63 Tweet
163 Takipçi
339 Takip Edilen


Alexey Nesvizhskii I think they tried (clumsily) to relate this instrument to a paper titled with the term "trueSCP". I felt that the use of this term in the paper was quite disdainful to other studies which paved the road to enable SCP.






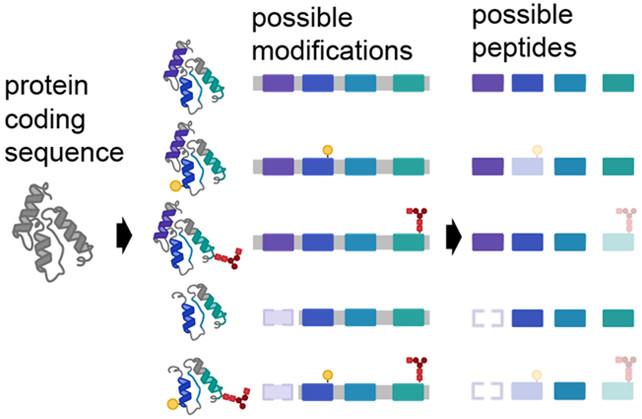
The @MacCossLab collaborated with MetaMorpheus @metamorpheus.bsky.social & others to reevaluate the current method of protein quantification in bottom-up proteomics. pubs.acs.org/doi/10.1021/ac…




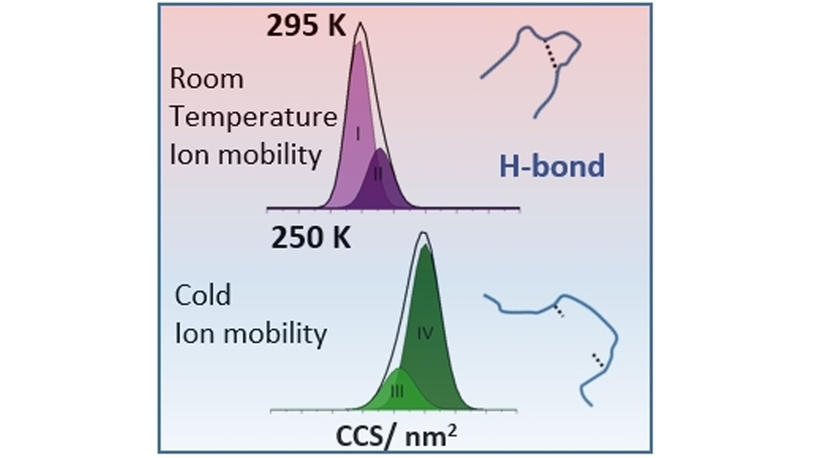
Cold Denaturation of Proteins in the Absence of Solvent: Implications for Protein Storage (Barran) @PerditaB Emma Norgate #openaccess onlinelibrary.wiley.com/doi/10.1002/an…



We have an outstanding lineup of speakers for the JPrOS Webinar "Top-Down Proteomics", gathering online from 🇯🇵, 🇩🇪, 🇫🇷 @[email protected], and 🇺🇸 Mowei Zhou, Joe Loo! Starts at 14:30 in continental Europe and bright and early at 5:30 on the West Coast! jproswebinar.com


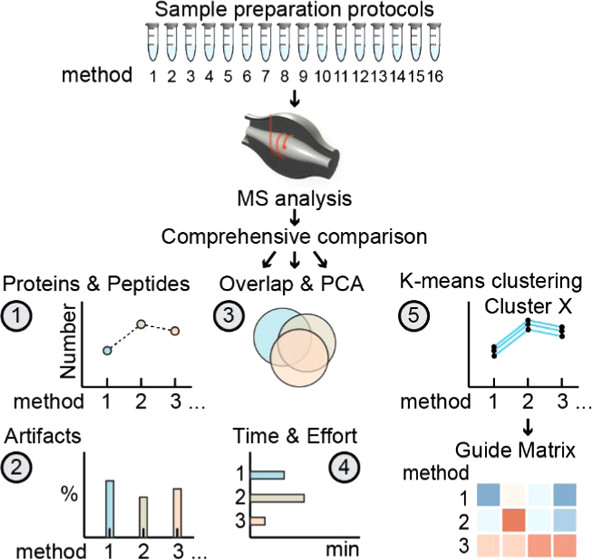
Searching for a universal, bottom-up proteomics method? Max Perutz Labs Vienna performed a comparative survey of the 16 most common methods to provide a basis for selection preparation strategies in MS proteomics. pubs.acs.org/doi/10.1021/ac…

Very happy to share our new paper published in Angewandte Chemie, where we investigated single C. elegans nematodes by top-down proteomics! #nanoproteomics #microfluidics onlinelibrary.wiley.com/doi/10.1002/an…

Thanks to #projectDEAL, our new top-down nanoproteomics paper published in Angewandte Chemie will remain Open Access :-)








