
Martin Schmitz
@martin_fschmitz
Ph.D. Student at the Genome Institute of Singapore and National University of Singapore
ID: 1519581435287719936
28-04-2022 07:37:20
9 Tweet
20 Followers
30 Following


After many months, I'm proud to share our work on untangling genome assembly graphs with GNNs. Or, GNNome Assembly (sorry not sorry)🧬 Paper: arxiv.org/abs/2206.00668 Code/Data: github.com/lvrcek/GNNome-… with Xavier Bresson, T. Laurent, Martin Schmitz, and Mile Sikic. Read more in 🧵
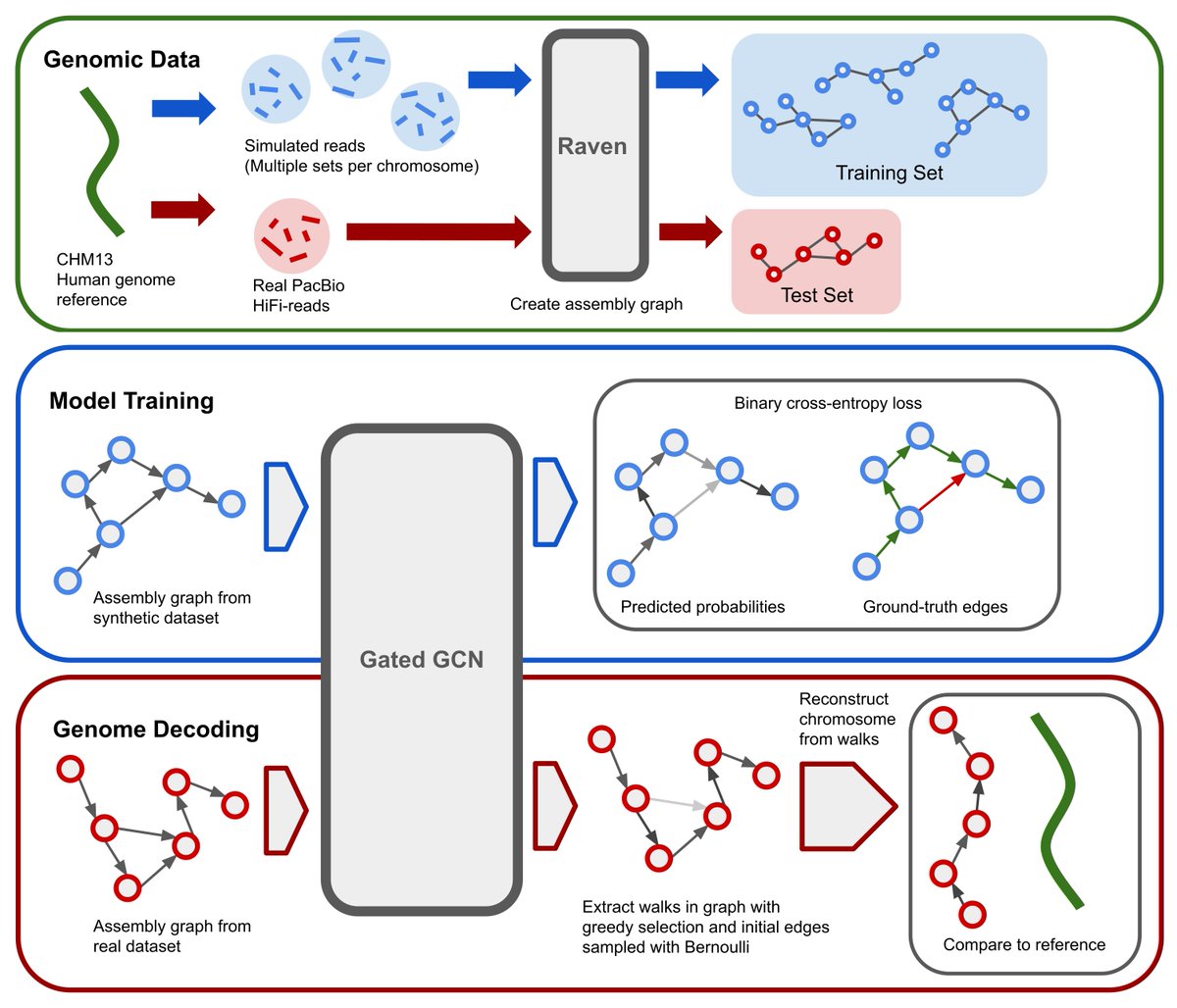

We present de novo genome assembly, based on Layout phase trained using GNNs, which outperforms the classical approach (removal of transitive edges, bubble popping, tips, unitigs,...) arxiv.org/abs/2206.00668 w/ Lovro Vrček Xavier Bresson T. Laurent, and Martin Schmitz


Long reads metagenome benchmark is out. Highlights. 1. In most cases Kraken, 2. minimap2/ram for slightly higher accuracy. 3. The right database is of huge importance 4. Check taxonomy files carefully. bmcbioinformatics.biomedcentral.com/articles/10.11… w/Niranjan Nagarajan Krešimir Križanović @jmaricb Sylvain




GNNome was published in Genome Research! This is a novel paradigm for de novo genome assembly based on GNNs. Without explicitly implementing any simplification strategies, it can achieve results comparable or higher than other SOTA tools. Paper, code, and overview are 👇 [1/8]
![Lovro Vrček (@lovrovrcek) on Twitter photo GNNome was published in <a href="/genomeresearch/">Genome Research</a>! This is a novel paradigm for de novo genome assembly based on GNNs. Without explicitly implementing any simplification strategies, it can achieve results comparable or higher than other SOTA tools. Paper, code, and overview are 👇 [1/8] GNNome was published in <a href="/genomeresearch/">Genome Research</a>! This is a novel paradigm for de novo genome assembly based on GNNs. Without explicitly implementing any simplification strategies, it can achieve results comparable or higher than other SOTA tools. Paper, code, and overview are 👇 [1/8]](https://pbs.twimg.com/media/GqAirreawAMb9Oy.jpg)
