Lukas Valihrach
@LukasValihrach
Group Leader | 🧬Single-Cell & Spatial Transcriptomics | Epigenomics | 🧠Neurobiology | Glial Cells in Health & Disease | @labgenex @IBT_CAS 🇪🇺 | ❤️ My Twins
ID:1199388618613280768
https://www.labgenexp.eu/ 26-11-2019 18:05:15
857 Tweets
2,6K Followers
777 Following







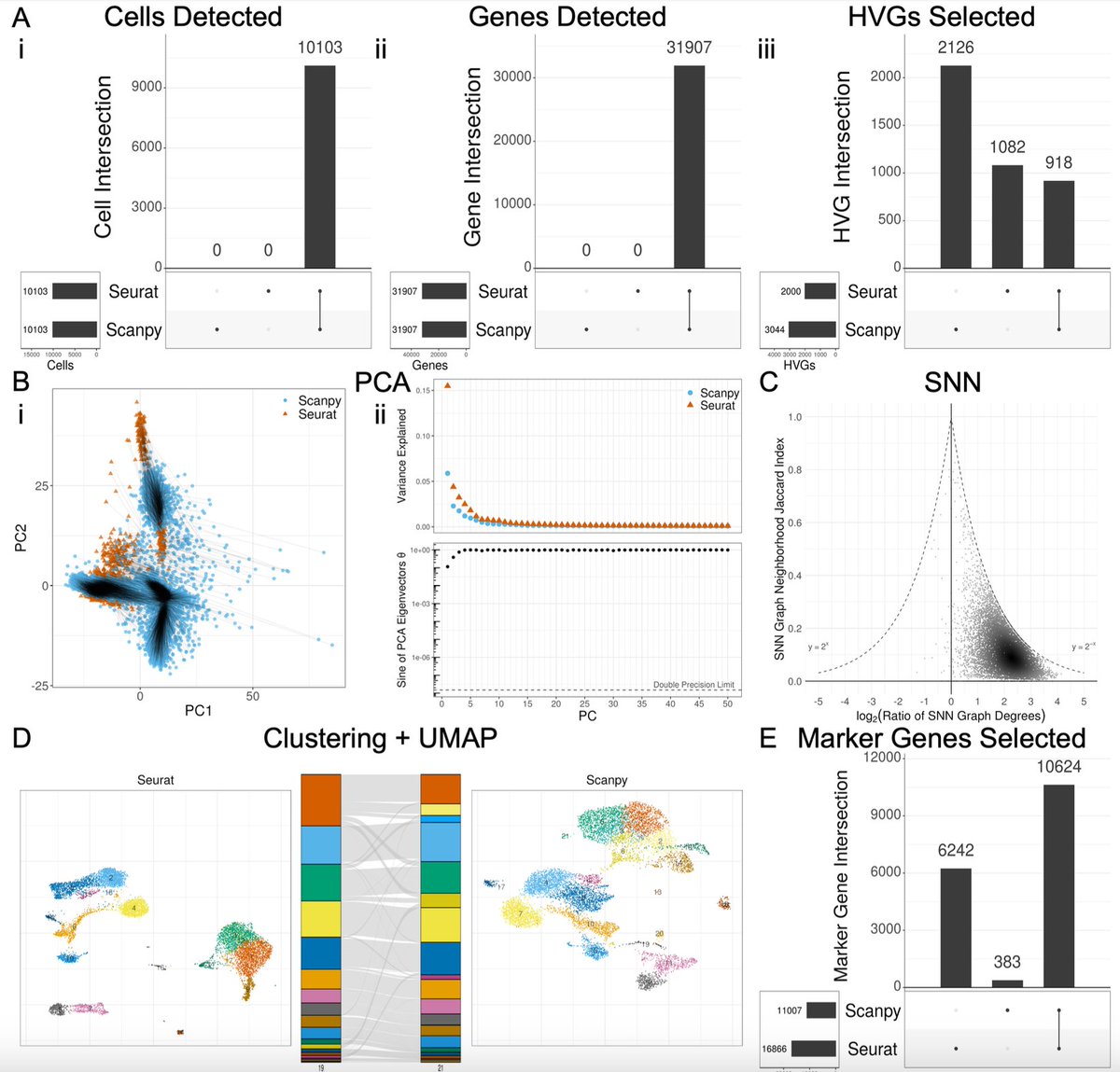
The choice of whether to use Seurat or Scanpy for single-cell RNA-seq analysis typically comes down to a preference of R vs. Python. But do they produce the same results? In biorxiv.org/content/10.110… w/ Joseph Rich et al. we take a close look. The results are 👀 1/🧵

In their Specious Art paper, Chari & Lior Pachter claim that tSNE/UMAP are as arbitrary as a random elephant shape. But are they?
We show in our comment that this is false and throws the tSNE/UMAP baby out with the bathwater!
Details in 🧵& paper:
biorxiv.org/content/10.110…
1/8

Just finished teaching another semester of genomic #datavisualization 🥳
Discover the awesome #dataviz made by students in the class for various #singlecell #spatialtranscriptomics data
Check out the course notes + #Rstats code to explore for yourself: jef.works/genomic-data-v…

PAPER ALERT!! Our latest work is out in Nature Neuroscience! We used humanised models to characterise in-depth the complex response of human microglia to Alzheimer's pathology, and we delved into the mechanisms and impact of genetics. nature.com/articles/s4159…



Don't miss our next #Bioinformatics seminar held by
Matematicko-fyzikální fakulta Univerzity Karlovy and Faculty of Science of Charles University. 'Revolutionizing Modern Biomedical Research with Single-Cell and Spatial Transcriptomics' by Lukas Valihrach from Institute of Biotechnology
📅 Wed 3. 4.
⏲️17:20
🏦Malostranske nam. 25
bioinformatics.cuni.cz/seminar


📢 How can we identify cell lineages from any scRNA-seq dataset ? Check our paper: nature.com/articles/s4146…. Very proud of massive amount of beautiful work by Almut Eisele, A. Dormann + fantastic collaboration with Marcel Tarbier and Pelechano lab. A 🧵
Wanna do standard 3’, 5’ etc 10x Genomics applications-seq without stress? No worries! FixNCut it baby!
Laura Jiménez Gracia YOU are the best! Holger Heyn Dr. Jasmine Plummer 👩🏻🔬🧬🇨🇦🇺🇸 CNAG Omniscope genomebiology.biomedcentral.com/articles/10.11…

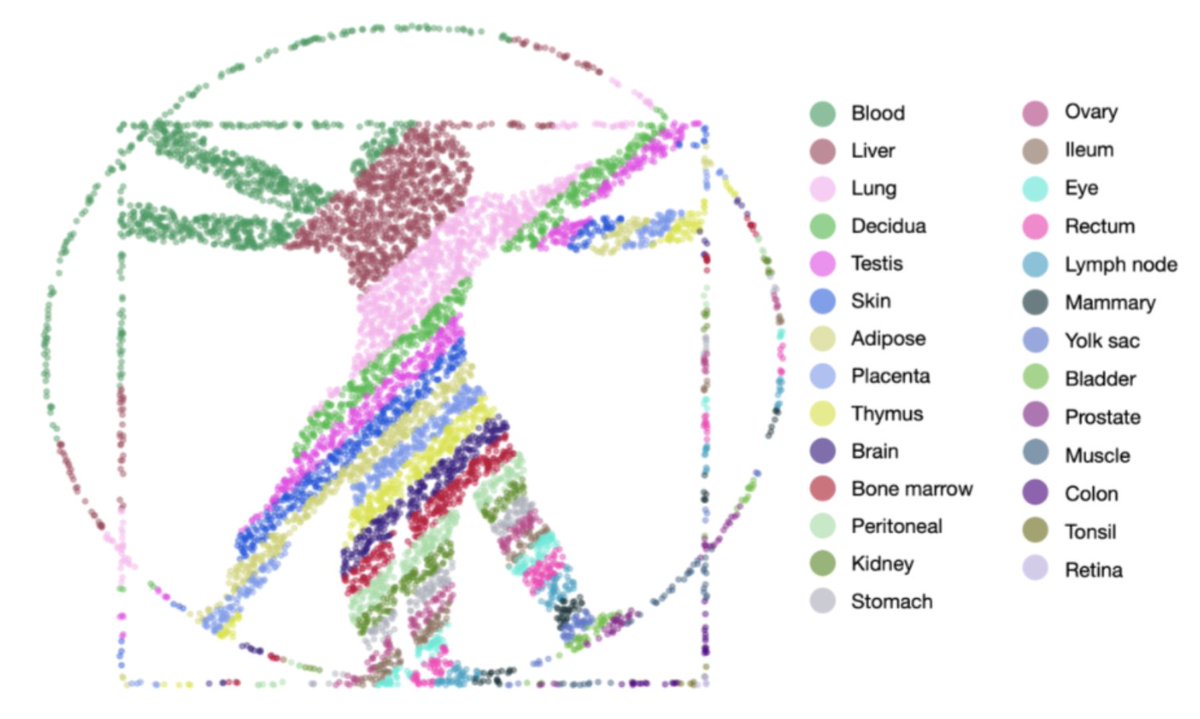
In a pair of bioRxiv preprints just posted w/ Sina Booeshaghi and Angel Galvez Merchan, we describe algorithms for an open data Commons Cell Atlas and demonstrate how a Human Commons Cell Atlas can be used for discovery.
biorxiv.org/content/10.110…
biorxiv.org/content/10.110…
1/🧵
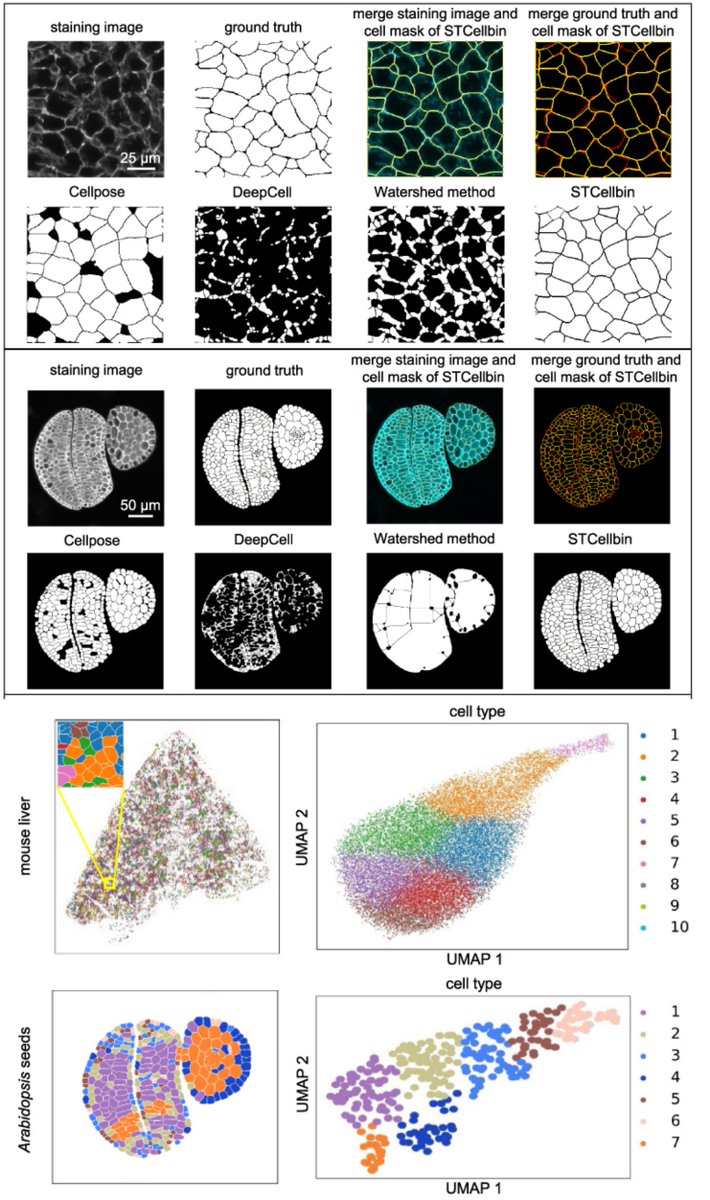
STCellbin
Use Nuclei+Cell Membrane/Wall stains to improve #CellSegmentation & single-cell-resolved #SpatialTranscriptomics
Tailored for #StereoSeq , but likely implementable for other ST methods
Susanne Brix & Xun Xu labs GigaByteJournal 2024
gigabytejournal.com/articles/110

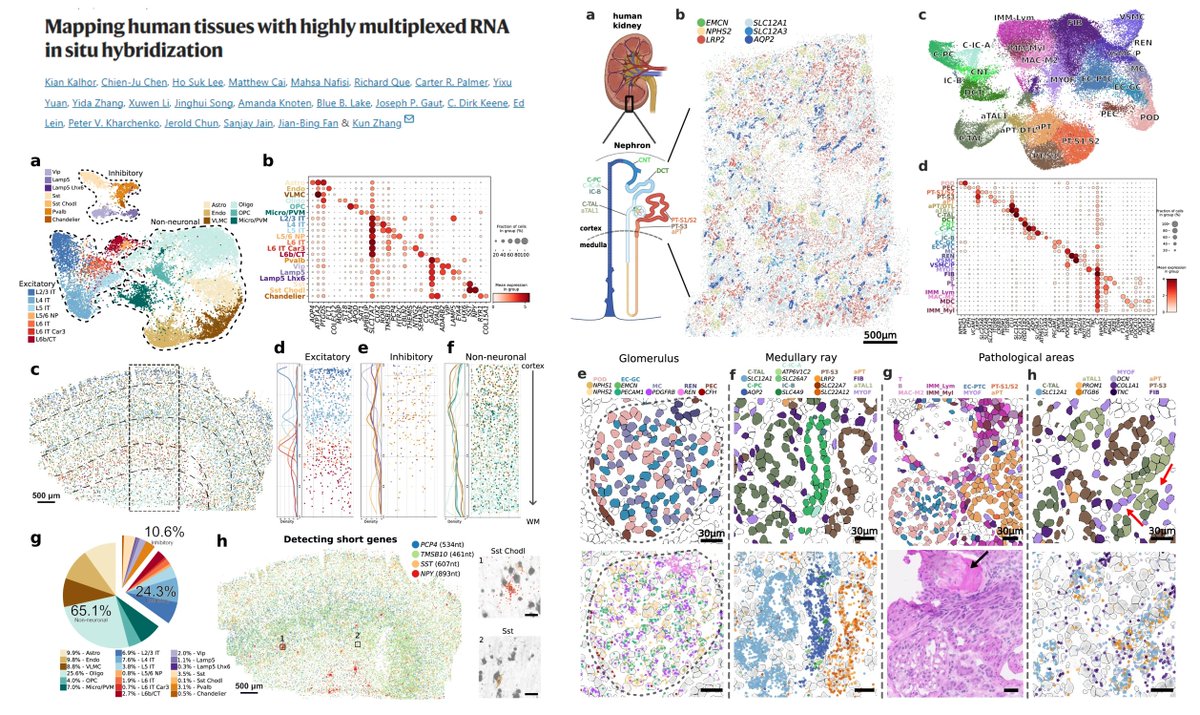
DART-FISH
#SpatialTranscriptomics
Pad-lock probe
Rolony
RiboSoma for cell segmentation
Detect <1kb short RNA (able to probe pri-microRNAs?😁)
Very nice reading if interested in how to design/optimize imaging-based ST methods👍
121 genes for human Cortex
300 genes for human…

What a PERFFect day to share our new preprint!
With Tsion Abay #BobStickels Meril Takizawa Ronan Chaligne (He/Him) and Ansu Satpathy, we introduce PERFF-seq, a new experimental approach to studying rare cells with scRNA-seq via transcript-specific enrichment. biorxiv.org/content/10.110… 1/n