Hannes Stärk
@hannesstaerk
@MIT PhD student • ML for molecules and biology bsky.app/profile/hannes…
ID: 1138012858988617728
http://hannes-stark.com 10-06-2019 09:19:43
2,2K Tweet
10,10K Takipçi
389 Takip Edilen


Nature Methods PNASNews 38/ Next paper in this series is #Boltz2 paper that described an open-source AI model that predicts both 3D protein structures and binding affinities — approaching the gold-standard FEP simulations, but 1000x faster ⚡ Why does this matter? Binding affinity = how tightly a





📌Notes on Boltz-2 Just watched the video talk led by Gabriele Corso Jeremy Wohlwend, and Saro Passaro that introduced Boltz-2, a structural biology foundation models. I summarized some learning notes below 🧵


Starkly Speaking tomorrow: Brandon Wood will present "UMA: A Family of Universal Models for Atoms" ai.meta.com/research/publi… Join us on Zoom 12pm ET / 6pm CEST: portal.valencelabs.com/starklyspeaking



Had a lot of fun learning diffusion and addressing key issues in protein diffusion with Giannis Daras Jeffrey Ouyang-Zhang TLDR: a few protein structure insights inspired us to design a new diffusion loss, training regime and dataset, resulting in significant performance improvements


Unfortunately, not able to attend #ICML2025 this year, but happy to share our accepted paper: 𝐒𝐏𝐇𝐈𝐍𝐗: 𝐒𝐭𝐫𝐮𝐜𝐭𝐮𝐫𝐚𝐥 𝐏𝐫𝐞𝐝𝐢𝐜𝐭𝐢𝐨𝐧 𝐮𝐬𝐢𝐧𝐠 𝐇𝐲𝐩𝐞𝐫𝐠𝐫𝐚𝐩𝐡 𝐈𝐧𝐟𝐞𝐫𝐞𝐧𝐜𝐞 𝐍𝐞𝐭𝐰𝐨𝐫𝐤 w/ Pietro Lio' arxiv.org/pdf/2410.03208 1/2

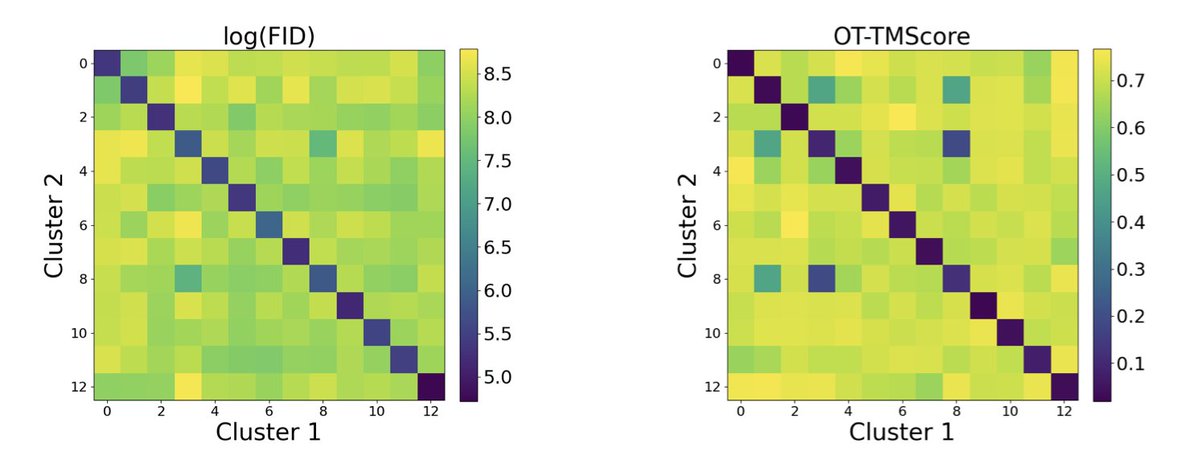
My labmate, officemate, and co-author Felix Faltings will be at #ICML2025 presenting ProxelGen in the GenBio Workshop @ ICML25 ! Anyone interested in discussing ProxelGen, ProtFID, or our current biomolecular design work? (ProxelGen link arxiv.org/pdf/2506.19820)

We also presented this work in Hannes Stärk's reading group. Go check it out if interested! (youtu.be/0r25eXy-Bgc?si…) Code coming soon! (12/12)


📢📢 "La-Proteina: Atomistic Protein Generation via Partially Latent Flow Matching" Fully atomistic. Partially latent. Structurally precise. Entirely generative. w/ Tomas Geffner*, Kieran Didi @ICLR*, et al. 📜 Project page & paper: research.nvidia.com/labs/genair/la… 🧵 Thread below... (1/n)

Excited to share: “Learning Diffusion Models with Flexible Representation Guidance” With my amazing coauthors Cai Zhou, Sharut Gupta, Johnson Lin, Stefanie Jegelka, Stephen Bates, Tommi Jaakkola Paper: arxiv.org/pdf/2507.08980 Code: github.com/ChenyuWang-Mon…