Adam Smiley
@adamsmiley17
BMBB PhD candidate at UMN in @WRgordonlab | @AndyArsham lab alumnus
ID: 1306685618748035074
17-09-2020 20:05:14
70 Tweet
71 Takipçi
321 Takip Edilen




Enzymes are promising catalysts for eco-friendly plastic recycling, but how can we make them faster? We developed a method to screen millions of enzyme mutants for enhanced plastic degradation! J. Am. Chem. Soc. 1/n doi.org/10.1021/jacs.3…

New paper out from my group pnas.org/doi/10.1073/pn…. CryoEM structure of the triplet polymerase ribozyme apoenzyme, with improved resolution from biorxiv.org/content/10.110…. Great collaboration with Ewan McRae and Ebbe Sloth Andersen

Check out our new generative AI tool for gene therapy published in Nature Machine Intelligence! Work conducted by the fantastic Suyue Lyu and Shahin Sowlati-Hashjin nature.com/articles/s4225…

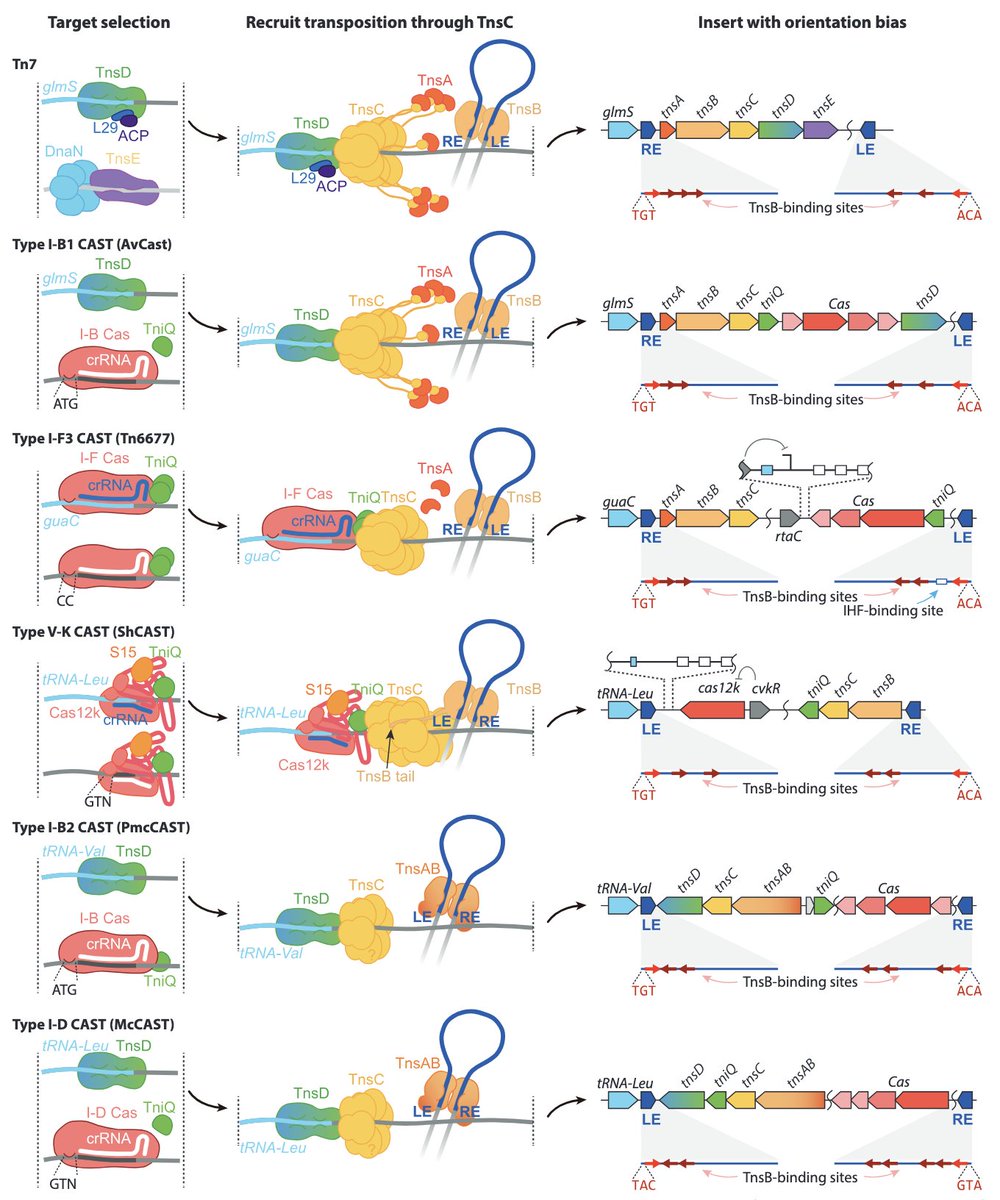
What if you could recombine any two DNA sequences in a programmable manner? We discovered an RNA-guided recombinase that can do just that! So happy to have co-led this project with Nicholas Perry


New Preprint, New Retrons! More than a hundred of them. We synthesized a ton of bioinformatically predicted retrons and put them through a battery of tests, from RT-DNA production to editing in human cells, E. coli, and phage biorxiv.org/content/10.110… Gladstone Institutes UC San Francisco UCSF Bioengineering & Therapeutic Sciences

New preprint alert! 🚨 Our latest work on plasmid mobility with @epcrocha & Amandine Nucci is now available. Identification of novel origins of transfer across bacterial plasmids. Read the full paper here: biorxiv.org/content/10.110… Thread🧵below!


Our manuscript "Biophysics-based protein language models for protein engineering" with Romero lab is now on bioRxiv. We present Mutational Effect Transfer Learning (METL), a protein language model trained on biophysical simulations, and showcase it for protein engineering. 1/



AmberBio is hiring: amber.bio/careers. I highly recommend looking into it if you want to work at a fast-moving startup with a creative and impactful technology. Basem Al-Shayeb, Ph.D. 🧬



Cellular repair of DNA breaks enables editing of the human genome. Do human cells repair RNA breaks, and can we use it to edit transcriptomes? Our work with Anna Nemudraia and Wiedenheft Lab is now online in Science Magazine! #ScienceResearch science.org/doi/10.1126/sc…


Now everyone customize/share protein language models for their custom task/dataset via Colaboratory 🤓 Paper: biorxiv.org/content/10.110… Colab: colab.research.google.com/drive/1nxYBed3… Credit: Jin Su, Zhikai Li, Chenchen Han, Yuyang Zhou, Junjie Shan, Xibin Zhou, Dacheng Ma, fajie yuan


We have trained ESM3 and we're excited to introduce EvolutionaryScale. ESM3 is a generative language model for programming biology. In experiments, we found ESM3 can simulate 500M years of evolution to generate new fluorescent proteins. Read more: evolutionaryscale.ai/blog/esm3-rele…